Molecular Dynamics Simulation
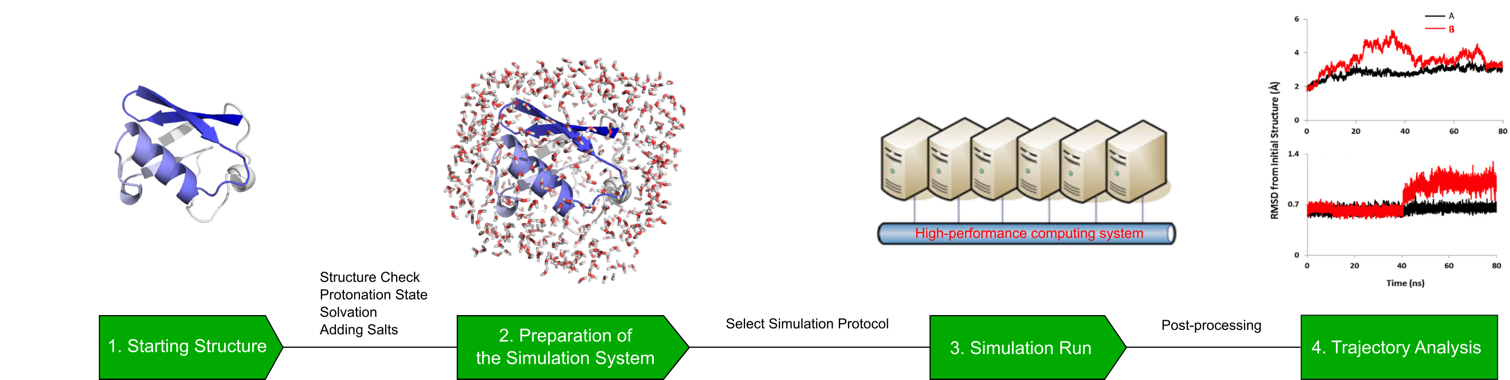
Molecular Dynamics Simulation Profacgen Molecular dynamics (md) is a computer simulation method for analyzing the physical movements of atoms and molecules. learn about the history, applications, methods, and limitations of md in chemical physics, materials science, and biophysics. The impact of molecular dynamics (md) simulations in molecular biology and drug discovery has expanded dramatically in recent years. these simulations capture the behavior of proteins and other biomolecules in full atomic detail and at very fine temporal resolution.

Molecular Dynamics Simulation Accuracy Speed Predictive Power Learn the basic idea, equations, properties, applications, and limitations of molecular dynamics simulation, a method to mimic the motion of atoms based on a potential energy function. see examples of md simulations of biomolecules, solvent effects, and drug binding. Md simulation mimics the changes in the structures of biological molecules over a given period of time, giving us atomic insights about the change in structure. this data helps us understand. Lammps is a classical molecular dynamics code with a focus on materials modeling. it can simulate atoms or particles at the atomic, meso, or continuum scale and supports parallel computing and accelerated performance. Molecular dynamics (md) simulation is a computational tool useful for predicting physical properties and elucidating reaction mechanisms at the atomic and molecular level.
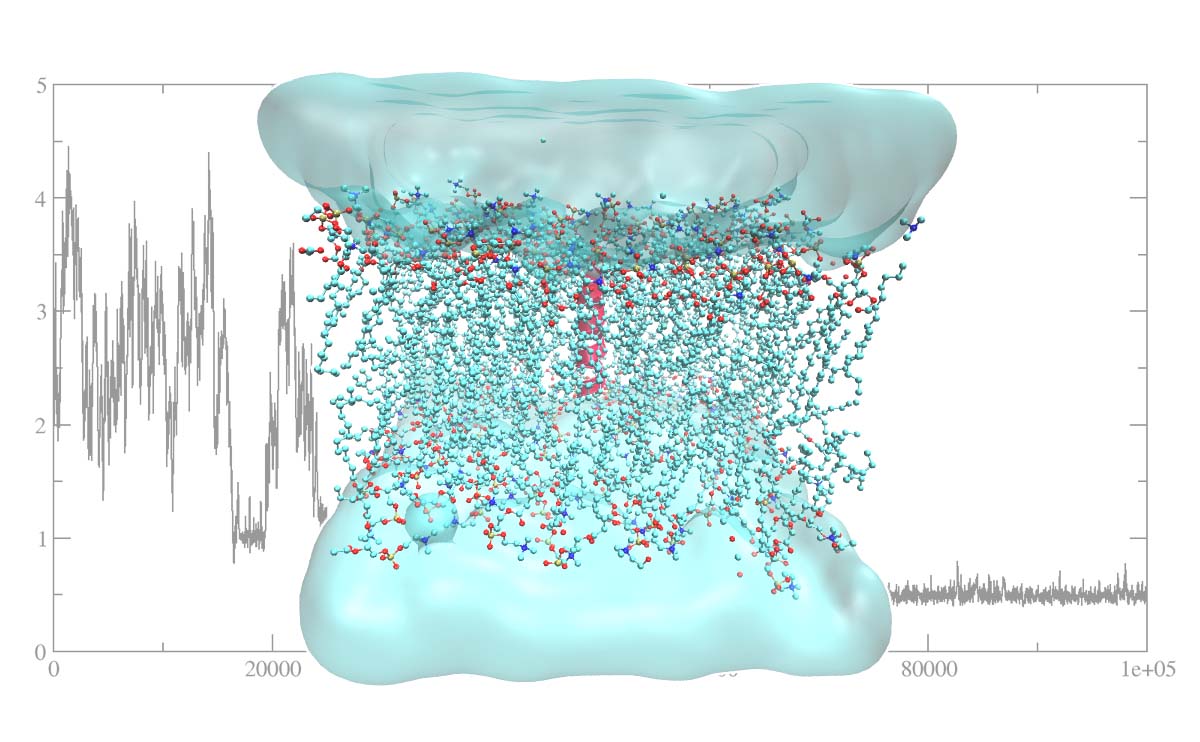
Getting Started With Molecular Dynamics Simulation Insilicosci Lammps is a classical molecular dynamics code with a focus on materials modeling. it can simulate atoms or particles at the atomic, meso, or continuum scale and supports parallel computing and accelerated performance. Molecular dynamics (md) simulation is a computational tool useful for predicting physical properties and elucidating reaction mechanisms at the atomic and molecular level. Learn how md simulations capture the behavior of biomolecules in full atomic detail and at very fine temporal resolution. see how md simulations are used in molecular biology and drug discovery, especially in neuroscience, and how they complement experimental structural biology techniques. Molecular dynamics simulation is a foundational computational approach for modeling the behavior of atoms and molecules in motion, enabling researchers to study the dynamics of molecular systems with up to millions of atoms using physical laws and mathematical models. Molecular dynamics (md) is a core methodology of molecular modeling and computational design for the study of the dynamics and temporal evolution of molecular systems. Molecular dynamics is a method that uses newton’s equations of motion to computationally simulate the time evolution of a set of interacting atoms. such techniques are dependent on a.

Molecular Dynamics Simulation Protac Learn how md simulations capture the behavior of biomolecules in full atomic detail and at very fine temporal resolution. see how md simulations are used in molecular biology and drug discovery, especially in neuroscience, and how they complement experimental structural biology techniques. Molecular dynamics simulation is a foundational computational approach for modeling the behavior of atoms and molecules in motion, enabling researchers to study the dynamics of molecular systems with up to millions of atoms using physical laws and mathematical models. Molecular dynamics (md) is a core methodology of molecular modeling and computational design for the study of the dynamics and temporal evolution of molecular systems. Molecular dynamics is a method that uses newton’s equations of motion to computationally simulate the time evolution of a set of interacting atoms. such techniques are dependent on a.
Comments are closed.