Introduction To Python Sequence To Function Analysis For Rna Rna

Rna Canada Arn Introduction To Python Sequence To Function Analysis The aim of this workshop is to show how molecular biology can be ‘recast’ into an ‘information centric’ framework, and for participants to develop essential python skills for bioinformatics. Quantification transcript quantification with salmon, post alignment qc with rseqc and dupradar, exploratory analysis and sample level qc (pca, sample distance heatmap) in python and r.

Introduction To Python Sequence To Function Analysis For Rna Rna Rna sequencing (rna seq) has transformed transcriptomic research, enabling researchers to perform large scale inspection of mrna levels in living cells. with the growing applicability of this technique to many scientific investigations, the analysis. "end to end rna seq data analysis with python based pipeline" is a comprehensive course designed to equip learners with the skills and knowledge necessary to conduct thorough rna seq data analysis using a python based pipeline. Among such methods, single cell rna seq (scrna seq) is having a profound impact on biology. here we introduce some of the key concepts of single cell rna seq technologies, with a focus on. Simply, various tools to work with rna data: sequences, alignments, structures, trajectories. if you want access the old version see the branch.
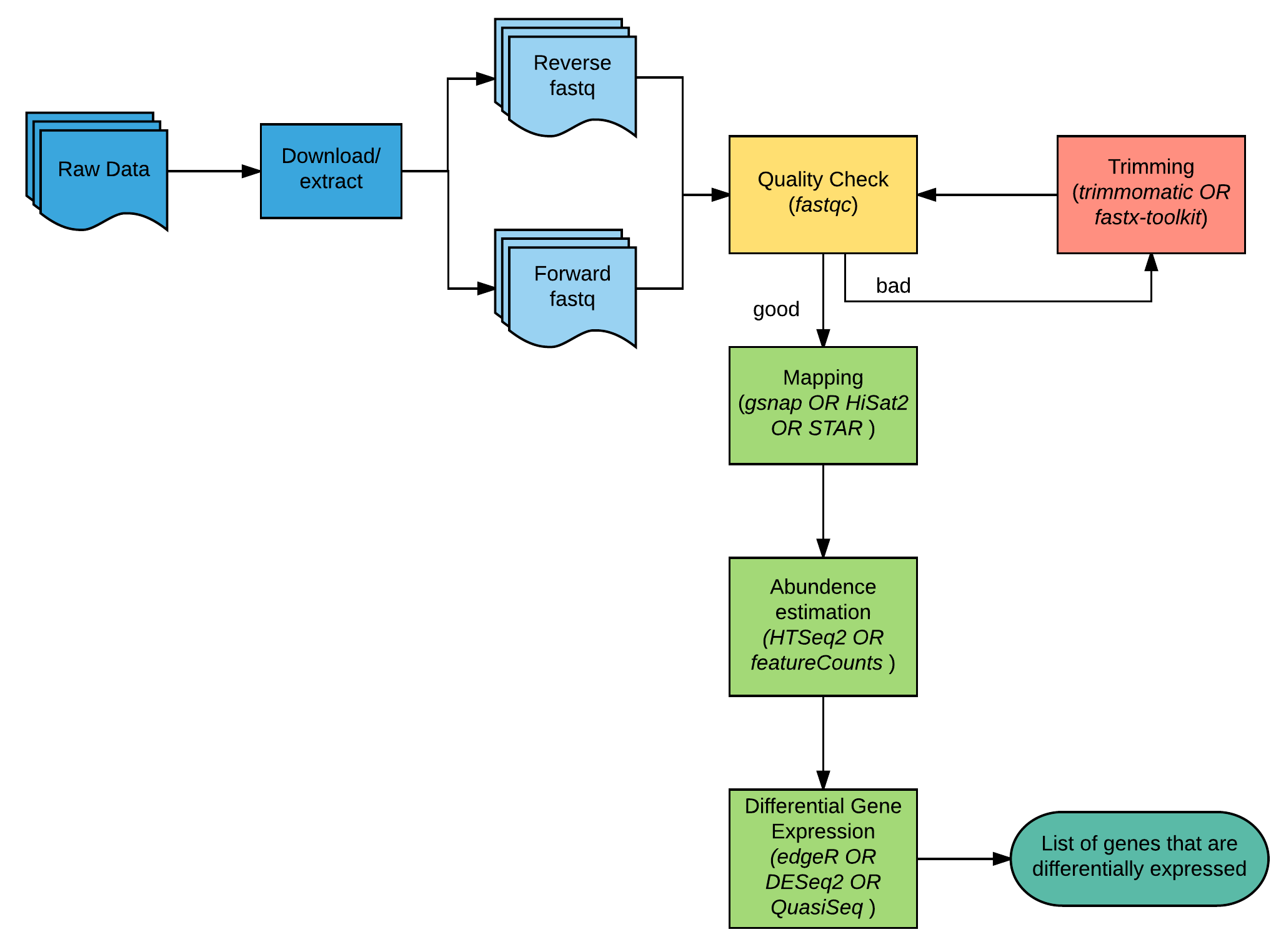
Rna Sequence Analysis Bioinformatics Workbook Among such methods, single cell rna seq (scrna seq) is having a profound impact on biology. here we introduce some of the key concepts of single cell rna seq technologies, with a focus on. Simply, various tools to work with rna data: sequences, alignments, structures, trajectories. if you want access the old version see the branch. Given a typical pipeline for rna sequencing (rna seq) data analysis consists of multiple steps, i divided it into three stages: (1) upstream data process; (2) midstream visualization of count matrix; and (3) downstream functional analysis. This document describes a suite of python tools for analysis of in vitro rna seq data (not intended for genomic analyses). the basic usage follows an object oriented model. We’re working with scanpy, the python iteration of the most widely used single cell toolkit. we’ve provided you with experimental data to analyse from a mouse dataset of fetal growth restriction bacon et al. 2018. In this chapter, we present an established pipeline for analyzing rna seq data, which involves a step by step flow starting from raw data obtained from a sequencer and culminating in the identification of differentially expressed genes with their functional characterization.

Introduction To Rna Sequencing And Analysis Rna Seq Blog Given a typical pipeline for rna sequencing (rna seq) data analysis consists of multiple steps, i divided it into three stages: (1) upstream data process; (2) midstream visualization of count matrix; and (3) downstream functional analysis. This document describes a suite of python tools for analysis of in vitro rna seq data (not intended for genomic analyses). the basic usage follows an object oriented model. We’re working with scanpy, the python iteration of the most widely used single cell toolkit. we’ve provided you with experimental data to analyse from a mouse dataset of fetal growth restriction bacon et al. 2018. In this chapter, we present an established pipeline for analyzing rna seq data, which involves a step by step flow starting from raw data obtained from a sequencer and culminating in the identification of differentially expressed genes with their functional characterization.
Comments are closed.