Introduction Cls Genomics Data

Introduction To Genomics Pdf Genome Gene This github site provides technical documentation and access information for researchers on the cls genomics data; more detailed information on the cohorts is available via the main cls website. Genotyped data and biological samples are available from all of cls’s adult cohort studies: the 1958 national child development study (ncds), the 1970 british cohort study (bcs70), next steps, and the millennium cohort study (mcs).

Introduction To Genomics Pdf This repository contains code to download documentation (pdfs) from cls's four established cohort studies ncds, bcs70, next steps, and mcs. the code places the pdfs into sub folders for each study sweep. Section 8.2 covers data applications via the cls dac for genetic data. the flowchart below shows the access procedure through the cls dac. polygenic indexes, proteomics and metabolomics are held by the uk data service under special licence. the data can be applied for directly via the ukds. Raw imputed genotype data – available for all cohort members who provided a dna sample that passed quality control. data access via the cls data access committe. The rich phenotyping of the cls cohorts, longitudinal repeat measure data, and their high participation rates make them highly suitable for both discovery and holdout prediction samples in gwas.

Introduction To Genomics Arthur M Lesk Z Library Pdf Raw imputed genotype data – available for all cohort members who provided a dna sample that passed quality control. data access via the cls data access committe. The rich phenotyping of the cls cohorts, longitudinal repeat measure data, and their high participation rates make them highly suitable for both discovery and holdout prediction samples in gwas. This tutorial uses the clc genomics workbench and clc single cell analysis module to reanalyze published data and recreate the results. you will learn to import different data types, perform qc and normalization, and using the flexible coloring and highlighting functionality of the umap plot. The genome build was updated to hg38 using liftover, which was implemented within the topmed server. imputed genotypes were then filtered with plink2.0alpha, excluding snps with an r2 info score < 0.8 and recoded as binary plink format. We have generated polygenic indexes (pgi) in all of the cls cohorts for multiple traits over a range of domains, as listed below. the pgi were generated through a standardised pipeline applied to the quality controlled and topmed imputed genetic data outlined on each cohort page of this site. Genotype calling was performed using genomestudio (v2.0, illumina) and quality control was completed using plink1.9 and plink2.0. 1681 samples were successfully read into genomestudio and mapped to a manifest file with the genome build grch38.

Introduction To Genomics This tutorial uses the clc genomics workbench and clc single cell analysis module to reanalyze published data and recreate the results. you will learn to import different data types, perform qc and normalization, and using the flexible coloring and highlighting functionality of the umap plot. The genome build was updated to hg38 using liftover, which was implemented within the topmed server. imputed genotypes were then filtered with plink2.0alpha, excluding snps with an r2 info score < 0.8 and recoded as binary plink format. We have generated polygenic indexes (pgi) in all of the cls cohorts for multiple traits over a range of domains, as listed below. the pgi were generated through a standardised pipeline applied to the quality controlled and topmed imputed genetic data outlined on each cohort page of this site. Genotype calling was performed using genomestudio (v2.0, illumina) and quality control was completed using plink1.9 and plink2.0. 1681 samples were successfully read into genomestudio and mapped to a manifest file with the genome build grch38.
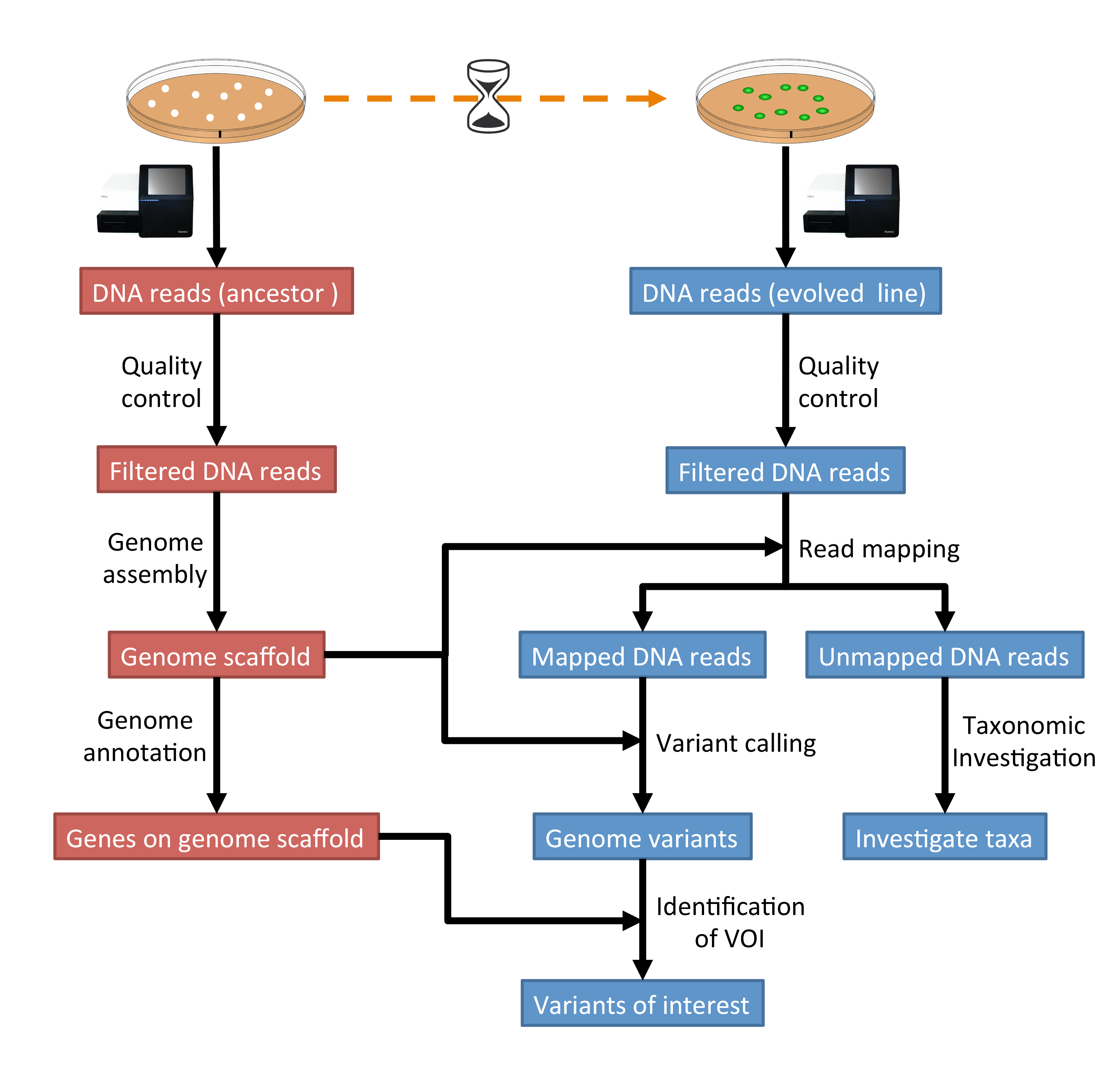
1 Introduction Genomics Tutorial 2020 2 0 Documentation We have generated polygenic indexes (pgi) in all of the cls cohorts for multiple traits over a range of domains, as listed below. the pgi were generated through a standardised pipeline applied to the quality controlled and topmed imputed genetic data outlined on each cohort page of this site. Genotype calling was performed using genomestudio (v2.0, illumina) and quality control was completed using plink1.9 and plink2.0. 1681 samples were successfully read into genomestudio and mapped to a manifest file with the genome build grch38.

1 Introduction Pdf Gene Genomics
Comments are closed.