Improve Speed And Accuracy Of Ngs Results For Viral Surveillance
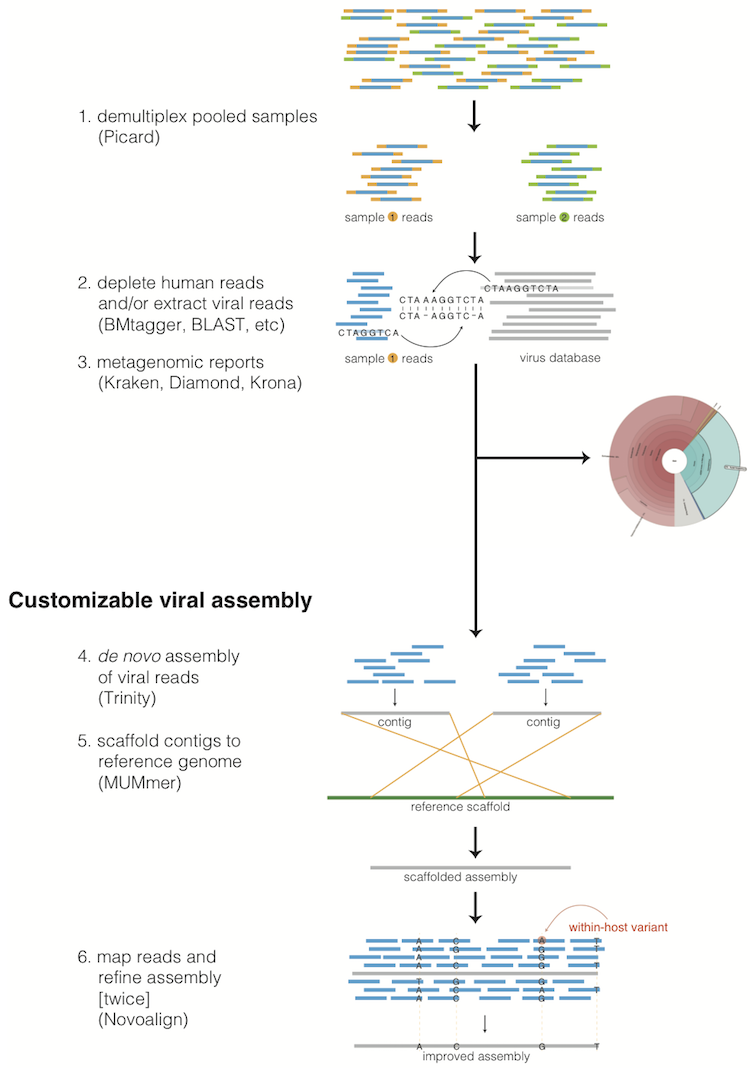
1 Description Of The Methods Viral Ngs V1 25 0 8 Ge144969e Documentation Collibri library prep kits for illumina systems enhance next generation sequencing (ngs) based viral surveillance with higher quality sequences and rapid protocols. understanding disease transmission pathways requires collaboration between governmental and private organizations. In order to maintain high quality next generation sequencing data, a variety of improvements at the levels of template preparation, sequencing strategy and data processing have been developed.

Pdf Surveillance Of Viral Emerging Infectious Diseases Using Ngs We developed a targeted next generation sequencing based variant surveillance strategy dubbed spike screen through sample pooling and selective sars cov 2 spike gene amplification in conjunction with cgs of individual cases to increase throughput and cost effectiveness. Summary: the amp published a consensus report on best practices for sars cov 2 genomic surveillance using next generation sequencing (ngs), providing critical guidelines to enhance data accuracy and public health outcomes. Cdc has launched national efforts to improve the ways sequencing data is used. these include efforts on a national viral genomics consortium to map sars cov 2 transmission and enhance newborn screening for precision public health. This study presents the development and validation of a genomic surveillance strategy using whole genome sequencing (wgs) on normalized pooled samples to detect and monitor sars cov 2 variants.
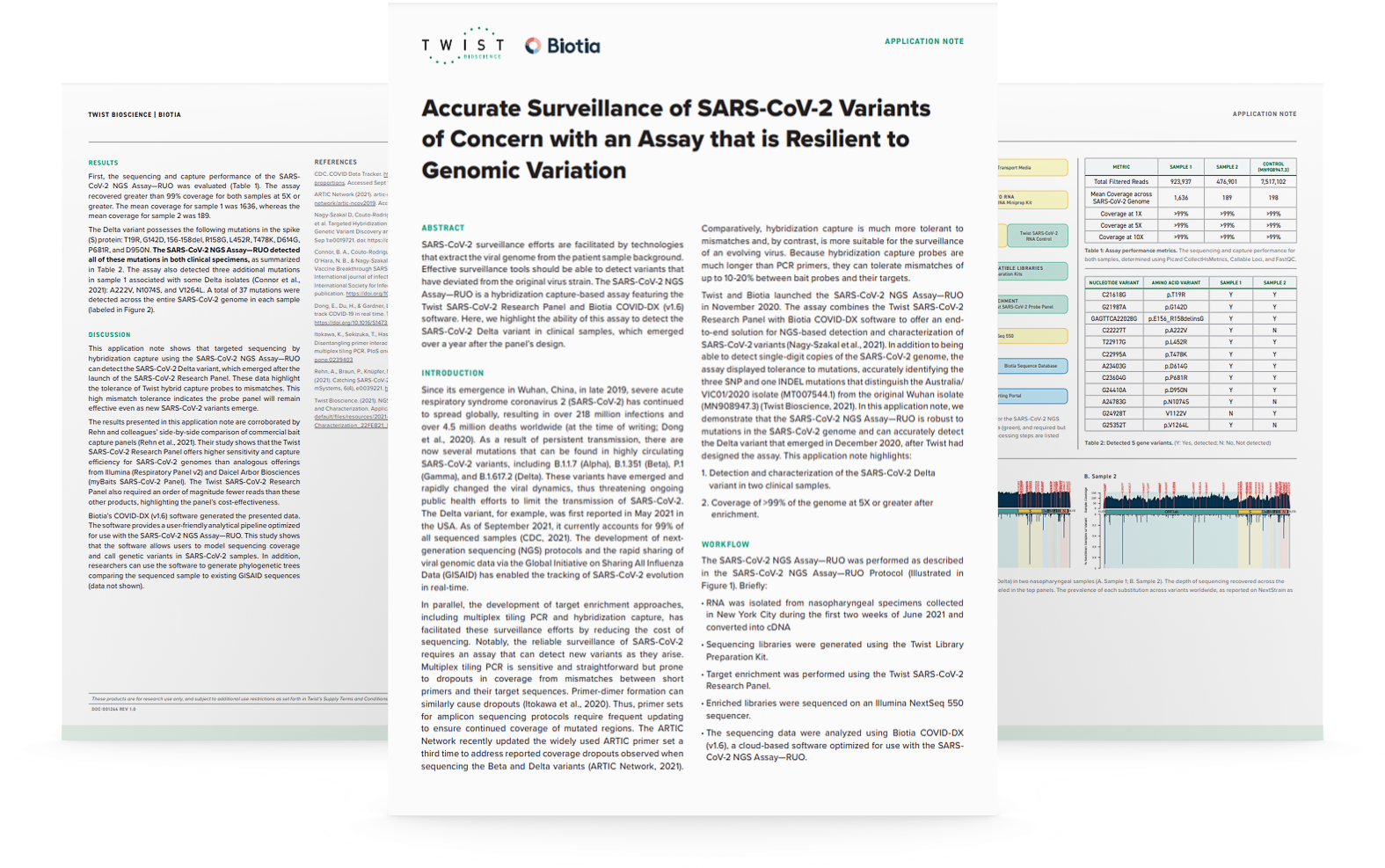

Achieve Highly Sensitive Viral Detection With The Sars Cov 2 Ngs Assay Cdc has launched national efforts to improve the ways sequencing data is used. these include efforts on a national viral genomics consortium to map sars cov 2 transmission and enhance newborn screening for precision public health. This study presents the development and validation of a genomic surveillance strategy using whole genome sequencing (wgs) on normalized pooled samples to detect and monitor sars cov 2 variants. Standardizing ngs methods, annotating nucleic acid sequences, and improving clinical documentation will ensure more reproducible results and support the design of cohesive pandemic surveillance in the future. The ability to respond quickly has relied on collaboration between diverse academic and public health professionals to increase sequencing capacity, develop methodologies to further improve the quality and efficiency of the data generated, and perform analyses required for genomic surveillance. The main goal of the virogenesis consortium was to develop new computational methods and software to overcome bioinformatics bottlenecks in the analysis and translation of ngs results for clinical and public health applications in the field of virology. In this paper, we present a fast and accurate workflow, virus identification workflow—viw, for virus identification and genome assembly based on ngs data to support viral pathogen research, especially during the covid 19 pandemic.

Implementing Fast Accessible Viral Surveillance With Nanopore Standardizing ngs methods, annotating nucleic acid sequences, and improving clinical documentation will ensure more reproducible results and support the design of cohesive pandemic surveillance in the future. The ability to respond quickly has relied on collaboration between diverse academic and public health professionals to increase sequencing capacity, develop methodologies to further improve the quality and efficiency of the data generated, and perform analyses required for genomic surveillance. The main goal of the virogenesis consortium was to develop new computational methods and software to overcome bioinformatics bottlenecks in the analysis and translation of ngs results for clinical and public health applications in the field of virology. In this paper, we present a fast and accurate workflow, virus identification workflow—viw, for virus identification and genome assembly based on ngs data to support viral pathogen research, especially during the covid 19 pandemic.

Local High Quality Ngs Can Improve Sars Cov 2 Surveillance Behind The main goal of the virogenesis consortium was to develop new computational methods and software to overcome bioinformatics bottlenecks in the analysis and translation of ngs results for clinical and public health applications in the field of virology. In this paper, we present a fast and accurate workflow, virus identification workflow—viw, for virus identification and genome assembly based on ngs data to support viral pathogen research, especially during the covid 19 pandemic.
Comments are closed.