Human Genome Sequence
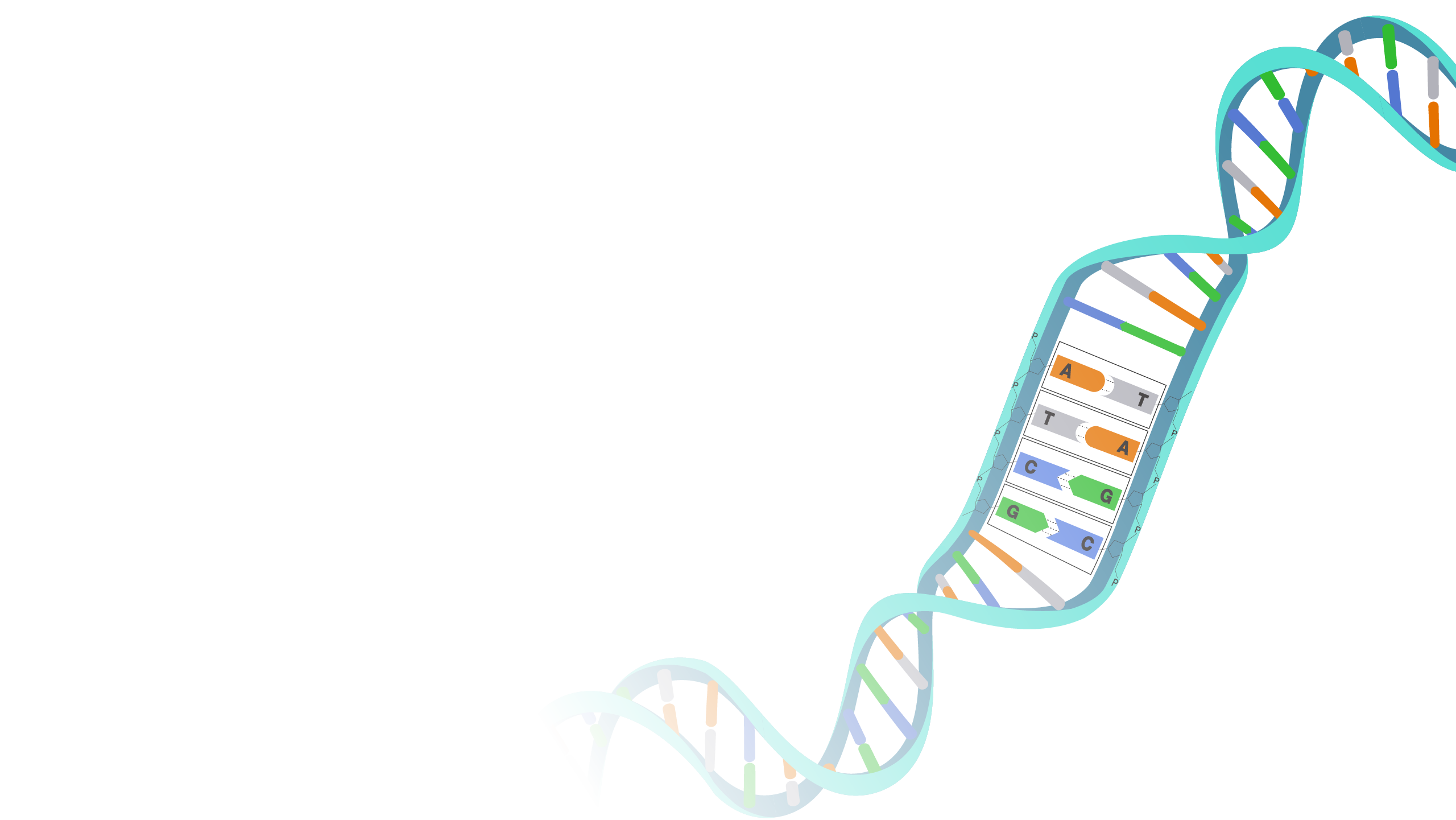

Human Genome Reference Sequence High accuracy long read sequencing has finally removed this technological barrier, enabling comprehensive studies of genomic variation across the entire human genome, which we expect to drive future discovery in human genomic health and disease. The human genome was the first of all vertebrates to be sequenced to such near completion, and as of 2018, the diploid genomes of over a million individual humans had been determined using next generation sequencing.
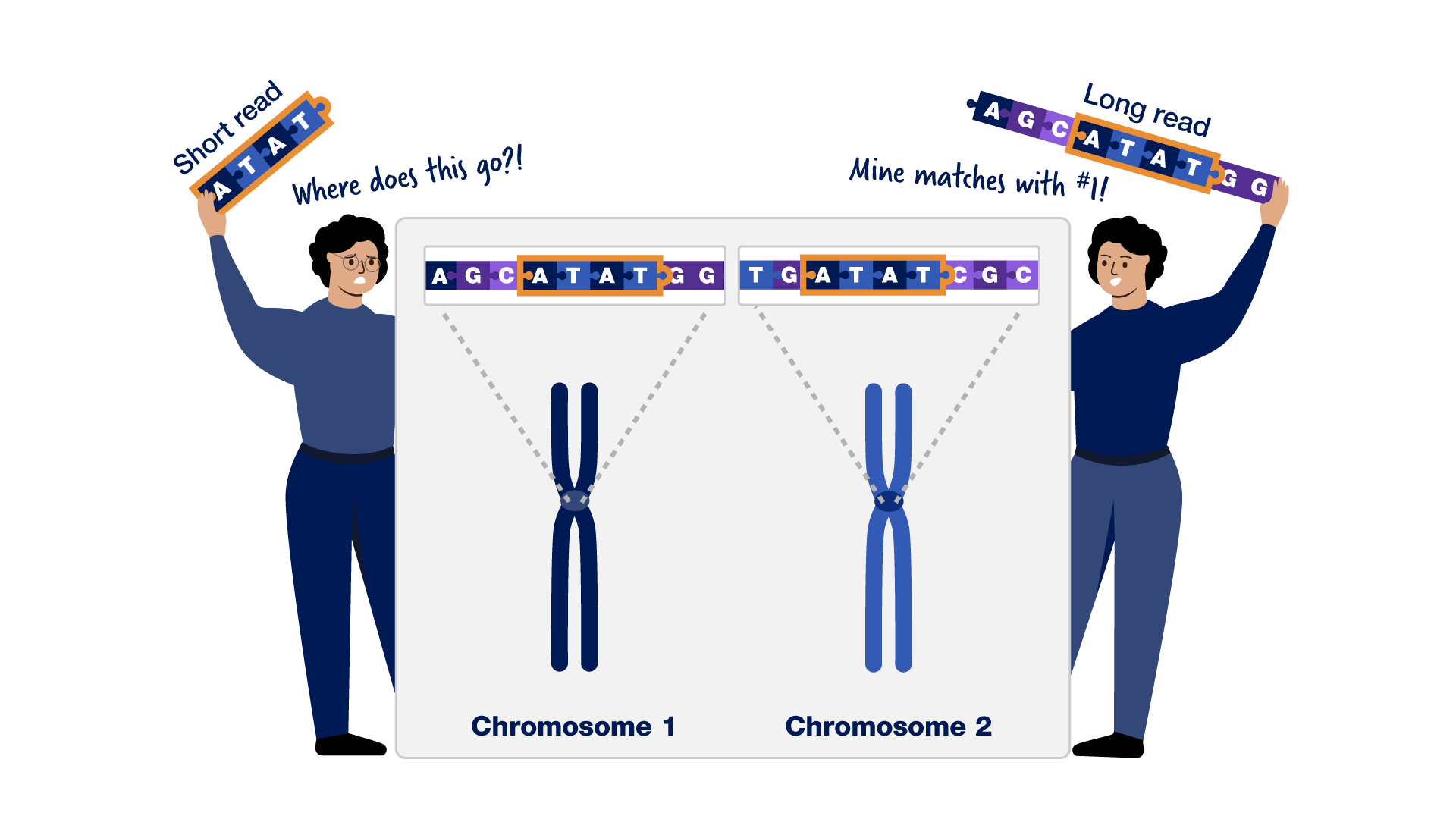
Completing The Human Genome Sequence The project was a voyage of biological discovery led by an international group of researchers looking to comprehensively study all of the dna (known as a genome) of a select set of organisms. Researchers finished sequencing the roughly 3 billion bases (or “letters”) of dna that make up a human genome. having a complete, gap free sequence of our dna is critical for understanding human genomic variation and the genetic contributions to certain diseases. Dna sequence comparisons between the consensus sequence and publicly funded genome data provided locations of 2.1 million single nucleotide polymorphisms (snps). a random pair of human haploid genomes differed at a rate of 1 bp per 1250 on average, but there was marked heterogeneity in the level of poly morphism across the genome. The human genome is not uniform. excepting identical (monozygous) twins, no two humans on earth share exactly the same genomic sequence. further, the human genome is not static. subtle and sometimes not so subtle changes arise with startling frequency.
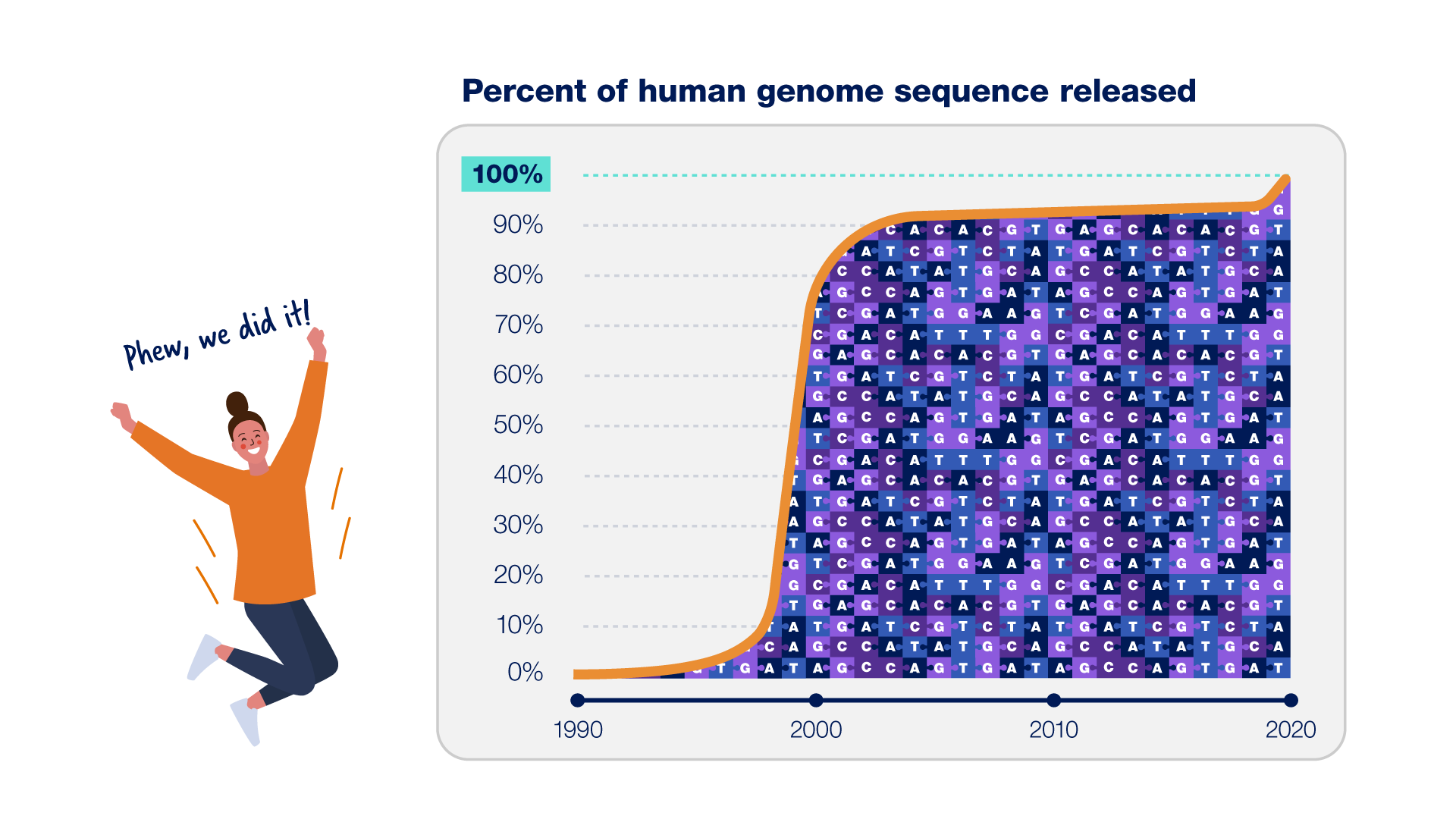
Completing The Human Genome Sequence Dna sequence comparisons between the consensus sequence and publicly funded genome data provided locations of 2.1 million single nucleotide polymorphisms (snps). a random pair of human haploid genomes differed at a rate of 1 bp per 1250 on average, but there was marked heterogeneity in the level of poly morphism across the genome. The human genome is not uniform. excepting identical (monozygous) twins, no two humans on earth share exactly the same genomic sequence. further, the human genome is not static. subtle and sometimes not so subtle changes arise with startling frequency. Addressing the remaining 8% of the genome, the telomere to telomere (t2t) consortium presents a complete 3.055 billion base pair sequence of a human genome, t2t chm13, that includes gapless. Twenty years after the initial drafts, a truly complete sequence of a human genome reveals what has been missing. since its initial release in 2000, the human reference genome has covered only the euchromatic fraction of the genome, leaving important heterochromatic regions unfinished. This week, the telomere to telomere (t2t) consortium has published a ‘ complete’ human genome sequence, filling in gaps that had stubbornly persisted for over 20 years since the publication of the original human genome project. Download dna sequence (fasta) convert your data to grch38 coordinates. display your data in ensembl. what can i find? protein coding and non coding genes, splice variants, cdna and protein sequences, non coding rnas. more about this genebuild. download fasta files for genes, cdnas, ncrna, proteins.
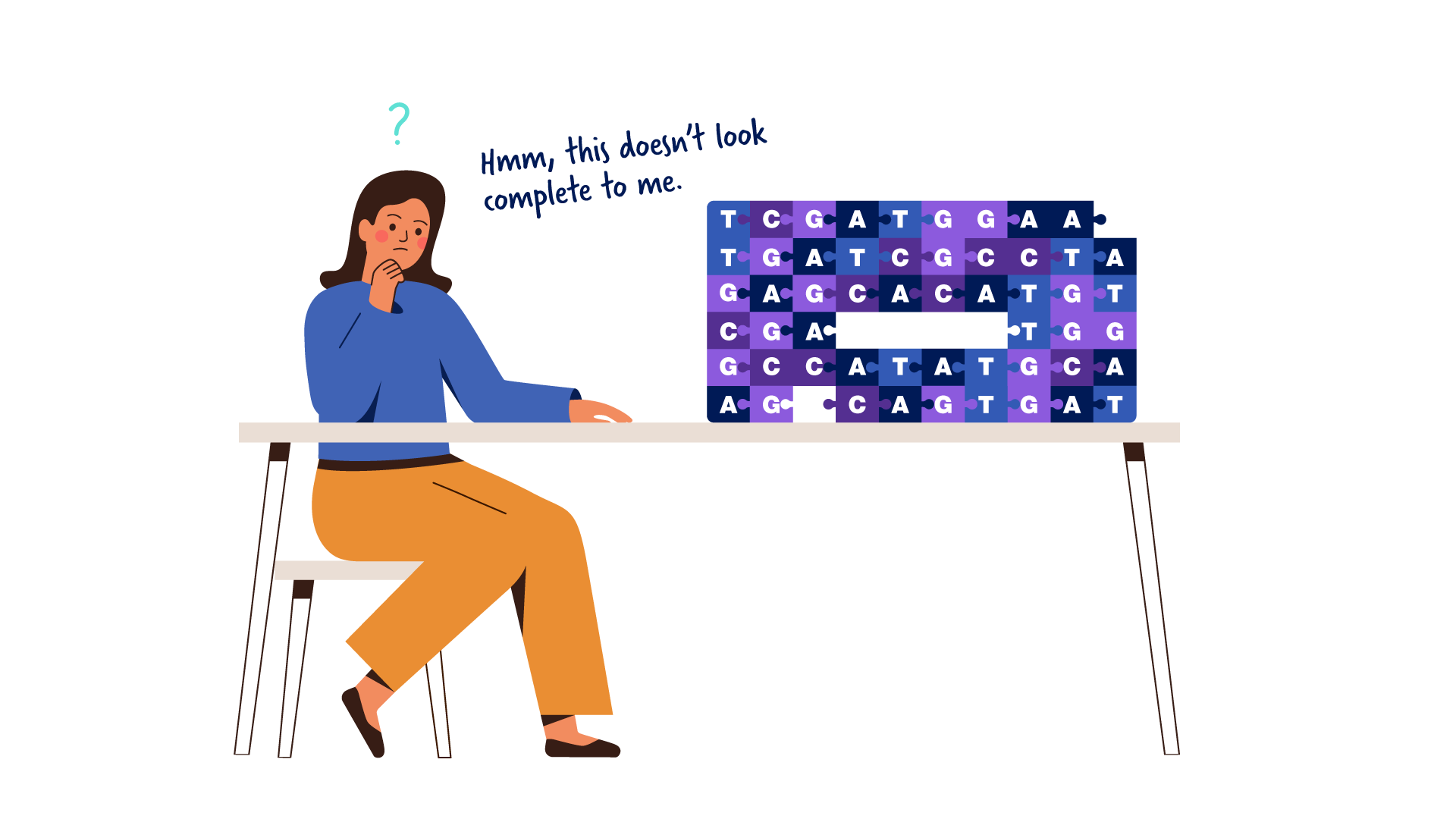

Completing The Human Genome Sequence Addressing the remaining 8% of the genome, the telomere to telomere (t2t) consortium presents a complete 3.055 billion base pair sequence of a human genome, t2t chm13, that includes gapless. Twenty years after the initial drafts, a truly complete sequence of a human genome reveals what has been missing. since its initial release in 2000, the human reference genome has covered only the euchromatic fraction of the genome, leaving important heterochromatic regions unfinished. This week, the telomere to telomere (t2t) consortium has published a ‘ complete’ human genome sequence, filling in gaps that had stubbornly persisted for over 20 years since the publication of the original human genome project. Download dna sequence (fasta) convert your data to grch38 coordinates. display your data in ensembl. what can i find? protein coding and non coding genes, splice variants, cdna and protein sequences, non coding rnas. more about this genebuild. download fasta files for genes, cdnas, ncrna, proteins.
Comments are closed.