How To Varseq Vcf Import The Golden Helix Blog
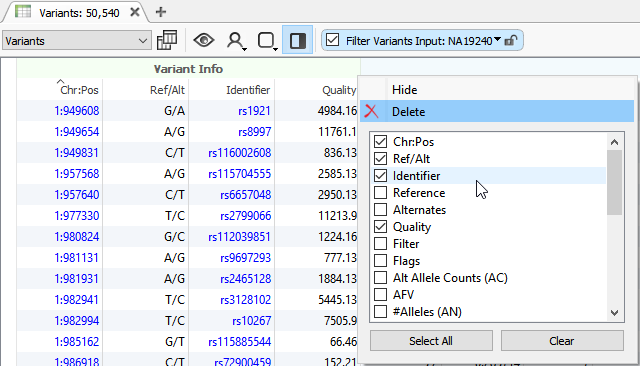
How To Varseq Vcf Import The Golden Helix Blog One major concern might be, how do i know varseq can handle importing my vcf? the purpose of this blog is exposing the readers to some key elements of varseq’s universal vcf importer. Varseq enables users to import structural variants for annotation, filtration, and subsequent clinical or other analyses. structural variants are often called during secondary analysis as belonging to two broad categories copy number variants (cnvs) with the file suffix " cnv.vcf" and breakends with the file suffix " sv.vcf".

How To Varseq Vcf Import The Golden Helix Blog It emphasizes the importance of accurate annotations and outlines the workflow from vcf files through to variant processing, highlighting various classification mechanisms and functional predictions. Our latest blog dives into the "how" and "why" behind this transformation, showcasing the versatility of varseq in handling diverse genetic data types. This blog will explain how you can automate the vcf and varseq project generation process that requires only a few commands. this will expedite your path to analysis as the newly created project will be ready to import rare variants quickly into vsclinical. In this webinar presentation, we will review workflow strategies for quality control and analysis of dna sequence variants using the varseq software package from golden helix.
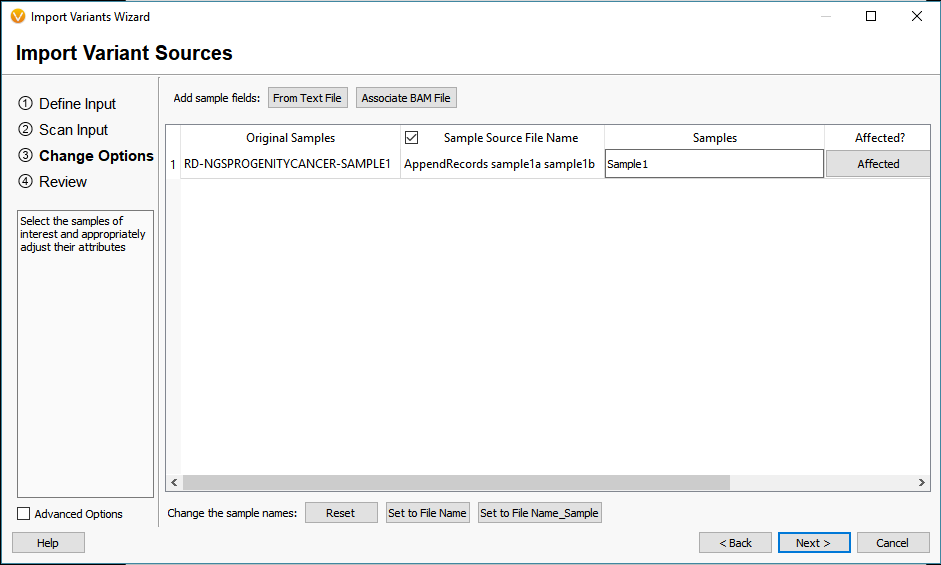
How To Varseq Vcf Import The Golden Helix Blog This blog will explain how you can automate the vcf and varseq project generation process that requires only a few commands. this will expedite your path to analysis as the newly created project will be ready to import rare variants quickly into vsclinical. In this webinar presentation, we will review workflow strategies for quality control and analysis of dna sequence variants using the varseq software package from golden helix. Vcf file format comes with a lot of interesting quality assurance and statistics fields that can be used for filtering in varseq. open your files in a text editor to see all the fields that are available in your files, each field will have a header line with a description of its content. Open the folder containing your vcf file in cursor. vcf files are massive text files that list every single mutation (variant) you have compared to the standard human reference genome. they are too big to open in a normal text editor, but they are incredibly easy to parse using python scripts. this is where cursor shines. The vcf2diem function processes calls for sites into the diem encoding for sites. the site order will be identical to that in the original vcf file (but see chapter sites not informative for genome polarisation). Explore common rna seq file formats: fastq, fasta, gtf gff, bam sam, bed, vcf. understand header structures, quality encoding, and how each format is used in the analysis pipeline.
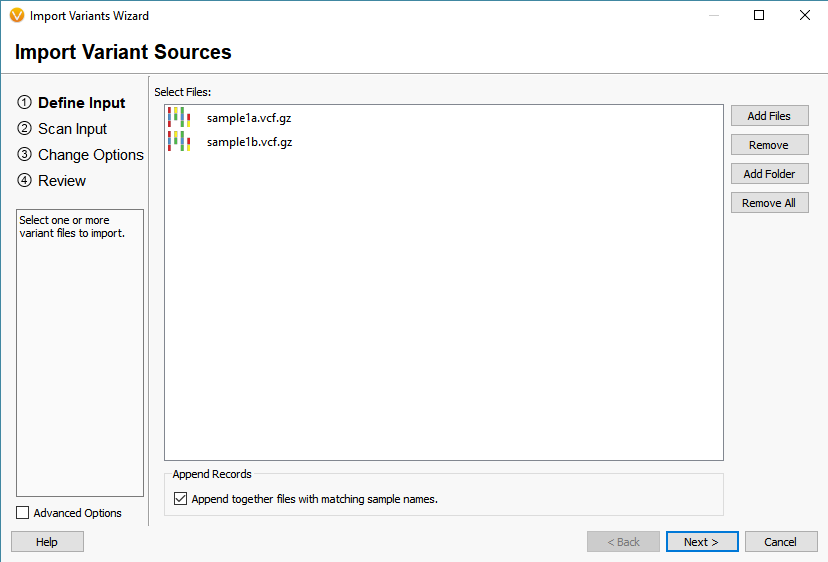
How To Varseq Vcf Import The Golden Helix Blog Vcf file format comes with a lot of interesting quality assurance and statistics fields that can be used for filtering in varseq. open your files in a text editor to see all the fields that are available in your files, each field will have a header line with a description of its content. Open the folder containing your vcf file in cursor. vcf files are massive text files that list every single mutation (variant) you have compared to the standard human reference genome. they are too big to open in a normal text editor, but they are incredibly easy to parse using python scripts. this is where cursor shines. The vcf2diem function processes calls for sites into the diem encoding for sites. the site order will be identical to that in the original vcf file (but see chapter sites not informative for genome polarisation). Explore common rna seq file formats: fastq, fasta, gtf gff, bam sam, bed, vcf. understand header structures, quality encoding, and how each format is used in the analysis pipeline.
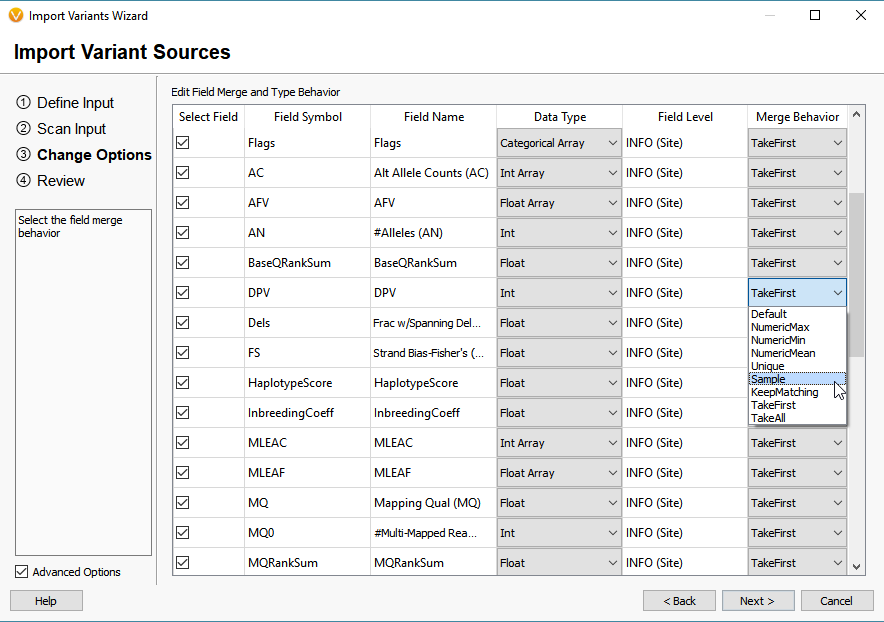
How To Varseq Vcf Import The Golden Helix Blog The vcf2diem function processes calls for sites into the diem encoding for sites. the site order will be identical to that in the original vcf file (but see chapter sites not informative for genome polarisation). Explore common rna seq file formats: fastq, fasta, gtf gff, bam sam, bed, vcf. understand header structures, quality encoding, and how each format is used in the analysis pipeline.

How To Varseq Vcf Import The Golden Helix Blog
Comments are closed.