High Throughput Technologies For Proteoform Detection Sheynkman Lab

Novel Mass Spectrometry Technologies For Proteoform Detection Learn more about the approaches we use in the sheynkman lab! © 2026 rector and visitors of the university of virginia. all rights reserved. At the sheynkman lab, we study disease relevant protein forms, or “proteoforms”, by integrating cutting edge analytical and computational approaches from genomi. a workflow for enhanced protein isoform detection through integration of long read rna seq and mass spectrometry based proteomics.
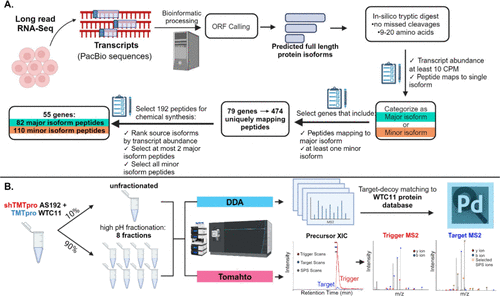
Novel Mass Spectrometry Technologies For Proteoform Detection This study introduces a systematic framework to estimate the explosion of proteoform diversity arising from dna, rna and ptm variation, highlighting the immense complexity of the human proteome. At the sheynkman lab, we study disease relevant protein forms, or “ proteoforms,” by integrating cutting edge analytical and computational approaches from genomics, proteomics, and systems biology. Recently, several new approaches for single molecule protein sequencing have emerged. with single aa resolution, these technologies are promising for distinguishing proteoforms by detecting subtle aa variations and ptms. In the sheynkman lab, we use bioinformatic tools to select the distinguishing regions of amino acids among proteoforms to use as target peptides for confirming their presence in samples.

Novel Mass Spectrometry Technologies For Proteoform Detection Recently, several new approaches for single molecule protein sequencing have emerged. with single aa resolution, these technologies are promising for distinguishing proteoforms by detecting subtle aa variations and ptms. In the sheynkman lab, we use bioinformatic tools to select the distinguishing regions of amino acids among proteoforms to use as target peptides for confirming their presence in samples. We introduce enhanced protein isoform detection through integrative “long read proteogenomics”. the core idea is to leverage long read rna seq to generate a sample specific database of full length protein isoforms. Interested in learning about our research? human disease genetics interactomics novel mass spectrometry technologies for proteoform detection next generation protein sequencing technology transcriptomics & long read proteogenomics. We introduce enhanced protein isoform detection through integrative “long read proteogenomics”. the core idea is to leverage long read rna seq to generate a sample specific database of full length protein isoforms. Quality control and filtering multiple quality control steps ensure high confidence results:.
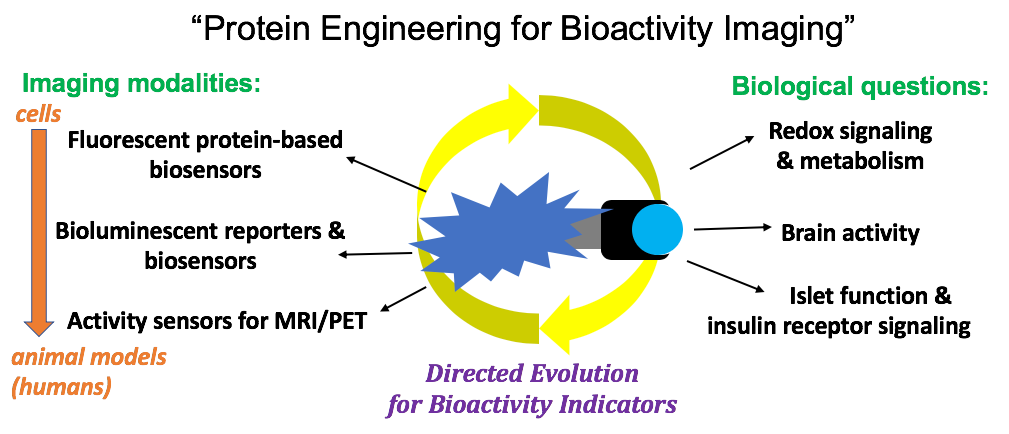
Research Sheynkman Lab We introduce enhanced protein isoform detection through integrative “long read proteogenomics”. the core idea is to leverage long read rna seq to generate a sample specific database of full length protein isoforms. Interested in learning about our research? human disease genetics interactomics novel mass spectrometry technologies for proteoform detection next generation protein sequencing technology transcriptomics & long read proteogenomics. We introduce enhanced protein isoform detection through integrative “long read proteogenomics”. the core idea is to leverage long read rna seq to generate a sample specific database of full length protein isoforms. Quality control and filtering multiple quality control steps ensure high confidence results:.

Sheynkman Lab Sheynkman Lab Website We introduce enhanced protein isoform detection through integrative “long read proteogenomics”. the core idea is to leverage long read rna seq to generate a sample specific database of full length protein isoforms. Quality control and filtering multiple quality control steps ensure high confidence results:.
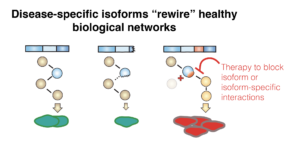
Sheynkman Lab Sheynkman Lab Website
Comments are closed.