High Resolution Spatial Transcriptomics From Histology Images Using
High Resolution Spatial Transcriptomics From Histology Images Using To address these limitations, we developed histosge, a method that employs a pathology image large model (pilm) to extract rich image features from histological images and utilizes a feature learning module to robustly generate high resolution gene expression profiles. Istar is presented, a method based on hierarchical image feature extraction that integrates st data and high resolution histology images to predict spatial gene expression with super resolution and enables gene expression prediction in tissue sections where only histology images are available.
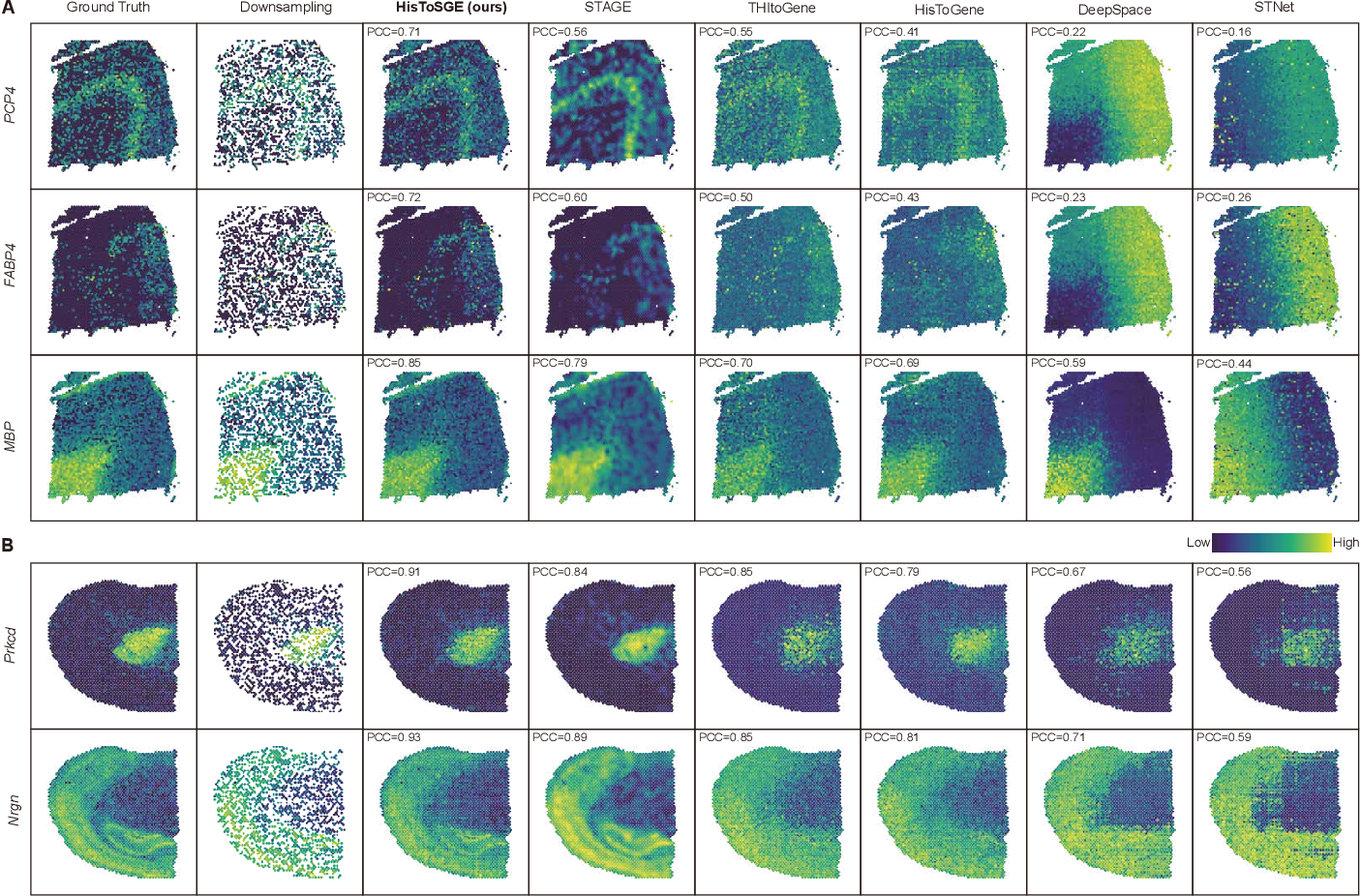
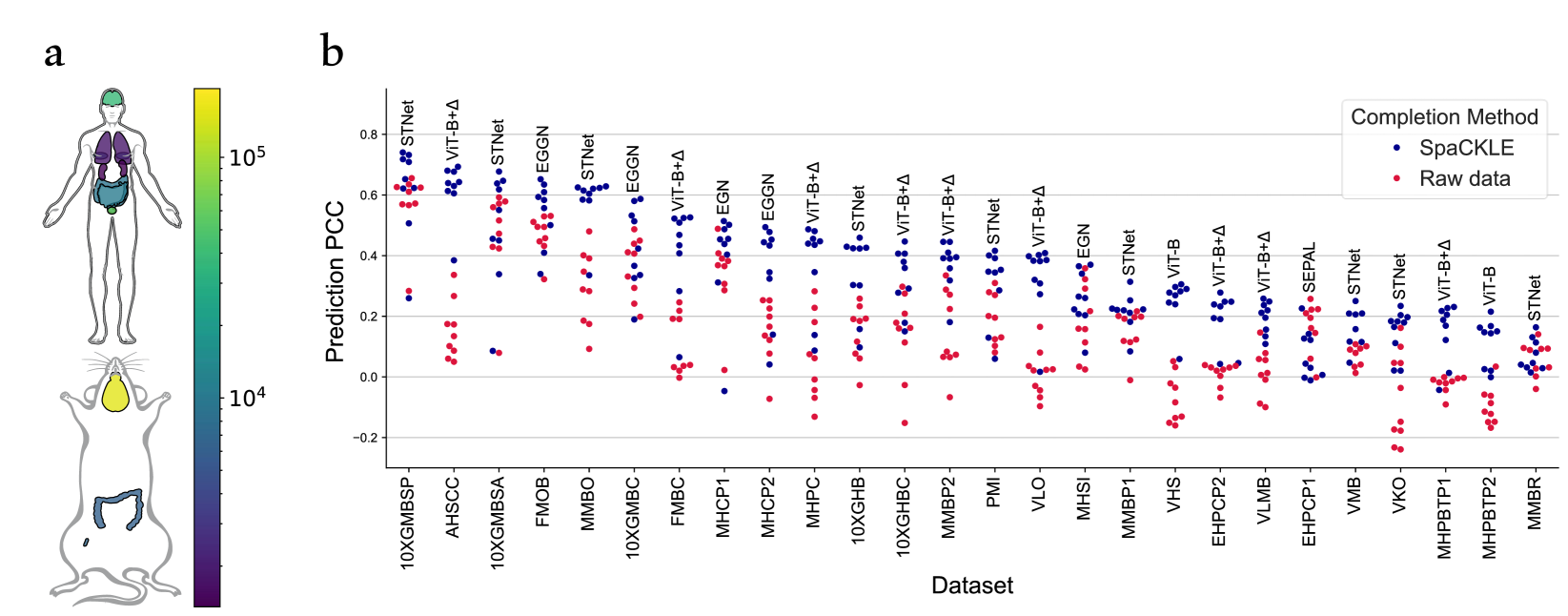
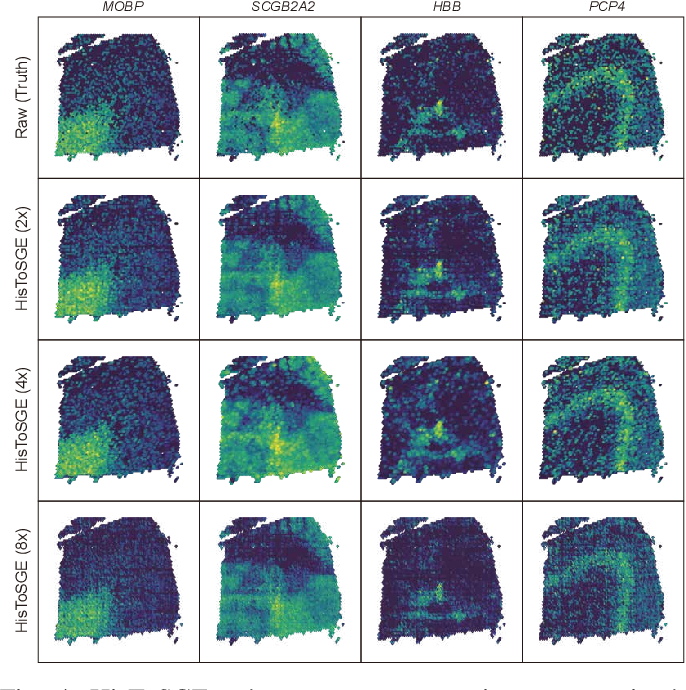
High Resolution Spatial Transcriptomics From Histology Images Using In this paper, we present spatial high definition embedding mapping (spahdmap), a multimodal dimension reduction framework to generate interpretable, high resolution embeddings of st data by. We evaluated histosge using four datasets and compared its performance with five existing methods. our results demonstrate that histosge can accurately generate high resolution gene expression profiles, enhance gene expression patterns, and preserve the original gene expression spatial structure. Histosge integrates histological image information, spatial information, and gene expression data, enhancing feature representation and generating high resolution profiles. Here we introduce a method that integrates spatial gene expression data with histological image data from the same tissue section to infer higher resolution expression maps.
High Resolution Spatial Transcriptomics From Histology Images Using Histosge integrates histological image information, spatial information, and gene expression data, enhancing feature representation and generating high resolution profiles. Here we introduce a method that integrates spatial gene expression data with histological image data from the same tissue section to infer higher resolution expression maps. We will leverage high resolution histological images combined with computer vision techniques such as cell segmentation and semantic tissue partitioning to identify potential single cell spatial locations and infer their corresponding gene expression profiles. Thor utilizes an anti shrinking markov diffusion method to infer single cell spatial transcriptomes from spot level data, effectively integrating cell morphology with spatial transcriptomics. the platform features 10 modules designed for cell level genomic and image analysis. To address these challenges, we introduce histex, a multimodal fusion approach that leverages a bidirectional cross attention mechanism and a general purpose foundation model. histex integrates spot based st data with histology images to predict super resolution (sr) spatial gene expression. Equipped with powerful foundation models pretrained on massive images, fmh2st employs a dual branch framework to integrate prior knowledge from foundation model and fine grained details from spot images.
Comments are closed.