Github Ncc Gap Gcatworkflow
Ncc Gap Github Contribute to ncc gap gcatworkflow development by creating an account on github. To generate sequence data and mutation data for use in the c cat utilization system, we have developed the genome analysis pipeline, g cat workflow ( github ncc gap gcatworkflow). g cat workflow is a genome analysis pipeline designed to run in grid engine computing environments.
Github Ncc Gap Ncc Ccat Gap Pages Url: github ncc gap gcatworkflow proper citation: gcat workflow (rrid:scr 027692) description: software cancer genome and rna sequencing data analysis pipeline that detects genomic variants and transcriptomic changes. 📅 last modified: wed, 02 nov 2022 09:42:05 gmt. Cwatch: ncc ccat gap gcatworkflow | none copyright © 2025 tz consulting gmbh & co. kg all rights reserved | | | | |. Contribute to ncc gap gcatworkflowcloud development by creating an account on github.

Github Ncc Gap Nanopore Workflow Scripts Cwatch: ncc ccat gap gcatworkflow | none copyright © 2025 tz consulting gmbh & co. kg all rights reserved | | | | |. Contribute to ncc gap gcatworkflowcloud development by creating an account on github. Gcat workflow rrid:scr 027692 github ncc gap gcatworkflow software cancer genome and rna sequencing data analysis pipeline that detects genomic variants and transcriptomic changes. To create sequence data and mutation data used in the c cat utilization system, we have previously developed the genome analysis pipeline called g cat workflow ( github ncc gap gcatworkflow). Specifically, we are progressing with the following initiatives: development and operation of a shared analysis environment for the genome raw data held by c cat (c cat calico). development of the cancer genome information analysis workflow, gcatworkflow. Ncc gap section of genome analysis platform, national cancer center pinned gcatworkflow public.
Github Ucaswangls Gap Ccot Gcat workflow rrid:scr 027692 github ncc gap gcatworkflow software cancer genome and rna sequencing data analysis pipeline that detects genomic variants and transcriptomic changes. To create sequence data and mutation data used in the c cat utilization system, we have previously developed the genome analysis pipeline called g cat workflow ( github ncc gap gcatworkflow). Specifically, we are progressing with the following initiatives: development and operation of a shared analysis environment for the genome raw data held by c cat (c cat calico). development of the cancer genome information analysis workflow, gcatworkflow. Ncc gap section of genome analysis platform, national cancer center pinned gcatworkflow public.
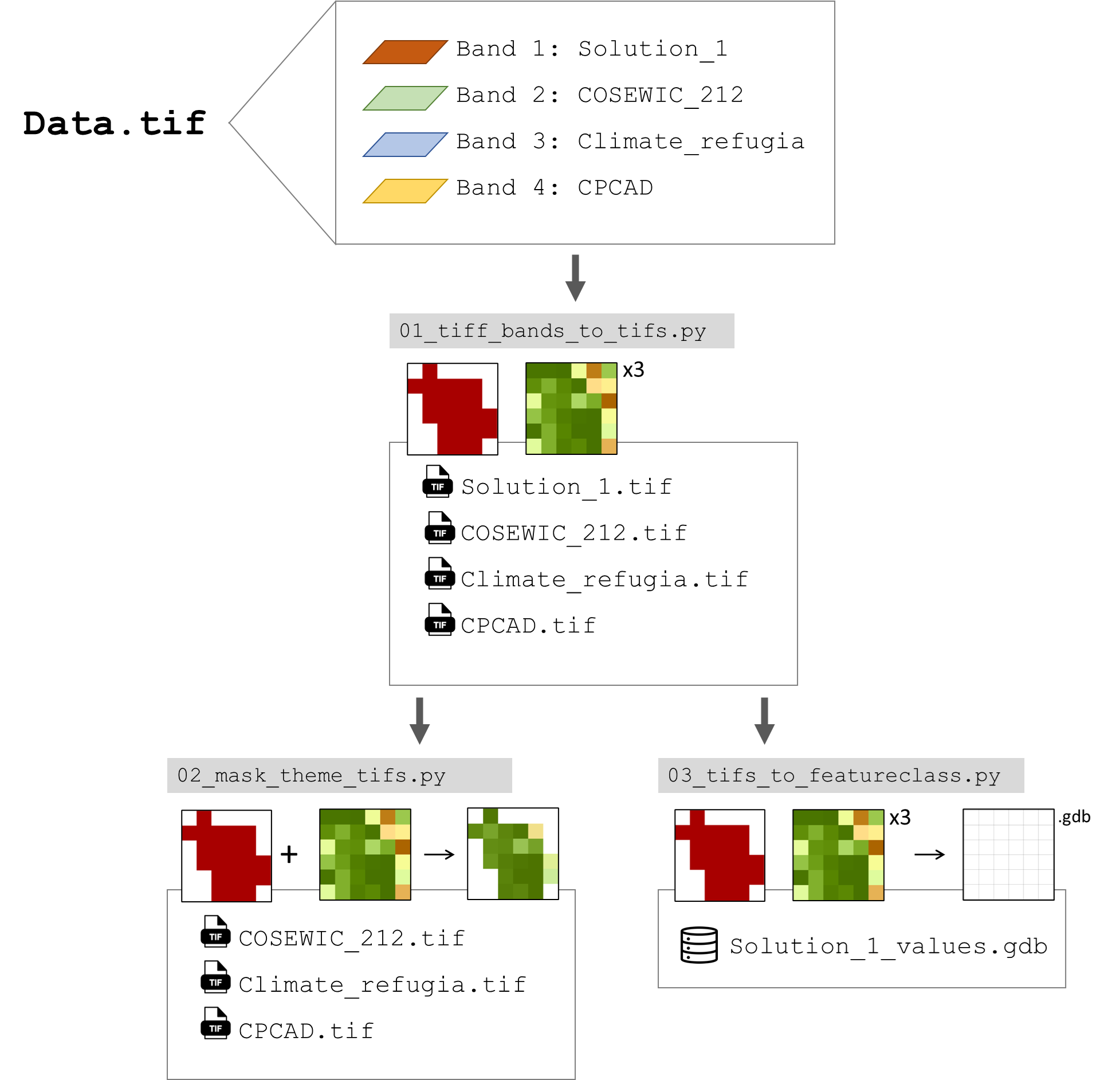
Github Ncc Cnc Wtw Output Processing Specifically, we are progressing with the following initiatives: development and operation of a shared analysis environment for the genome raw data held by c cat (c cat calico). development of the cancer genome information analysis workflow, gcatworkflow. Ncc gap section of genome analysis platform, national cancer center pinned gcatworkflow public.
Github Lazyhacker Gapbuffer C Implementation Of A Gap Buffer
Comments are closed.