Github Jiangbiolab Deepst Identify Spatial Domain
Github Jiangbiolab Deepst Identify Spatial Domain The output of deepst can be applied to identify spatial domains, batch effect correction and downstream analysis. In this paper, we propose deepst, a deep learning framework that integrates spatial location, histology, and gene expression to model spatially embedded representations to identify spatial domains with similar expression and histology.
Github Jiangbiolab Deepst Identify Spatial Domain Here, we present deepst, an accurate and universal deep learning framework to identify spatial domains, which performs better than the existing state of the art methods on benchmarking datasets of the human dorsolateral prefrontal cortex. Here, we present deepst, an accurate and universal deep learning framework to identify spatial domains, which performs better than the existing state of the art methods on benchmarking datasets of the human dorsolateral prefrontal cortex. By integrating gene expression, spatial information, and optionally histology images, deepst can identify biologically meaningful spatial domains in various tissue types. In this paper, we propose deepst, a deep learning framework that integrates spatial location, histology, and gene expression to model spatially embedded representations to identify spatial domains with similar expression and histology.
关于merfish数据的部分 Issue 8 Jiangbiolab Deepst Github By integrating gene expression, spatial information, and optionally histology images, deepst can identify biologically meaningful spatial domains in various tissue types. In this paper, we propose deepst, a deep learning framework that integrates spatial location, histology, and gene expression to model spatially embedded representations to identify spatial domains with similar expression and histology. Here, we present deepst, an accurate and universal deep learning framework to identify spatial domains, which performs better than the existing state of the art methods on benchmarking. To obtain a sufficiently flexible model, we developed deepst, a graph convolutional network that learns on graphs defined from spatial transcriptomics data sets. Deepst first uses h&e staining to extract tissue morphology information through a pre trained deep learning model, and normalizes each spot’s gene expression according to the similarity of adjacent spots.
Keyerror Use Quality Issue 11 Jiangbiolab Deepst Github Here, we present deepst, an accurate and universal deep learning framework to identify spatial domains, which performs better than the existing state of the art methods on benchmarking. To obtain a sufficiently flexible model, we developed deepst, a graph convolutional network that learns on graphs defined from spatial transcriptomics data sets. Deepst first uses h&e staining to extract tissue morphology information through a pre trained deep learning model, and normalizes each spot’s gene expression according to the similarity of adjacent spots.
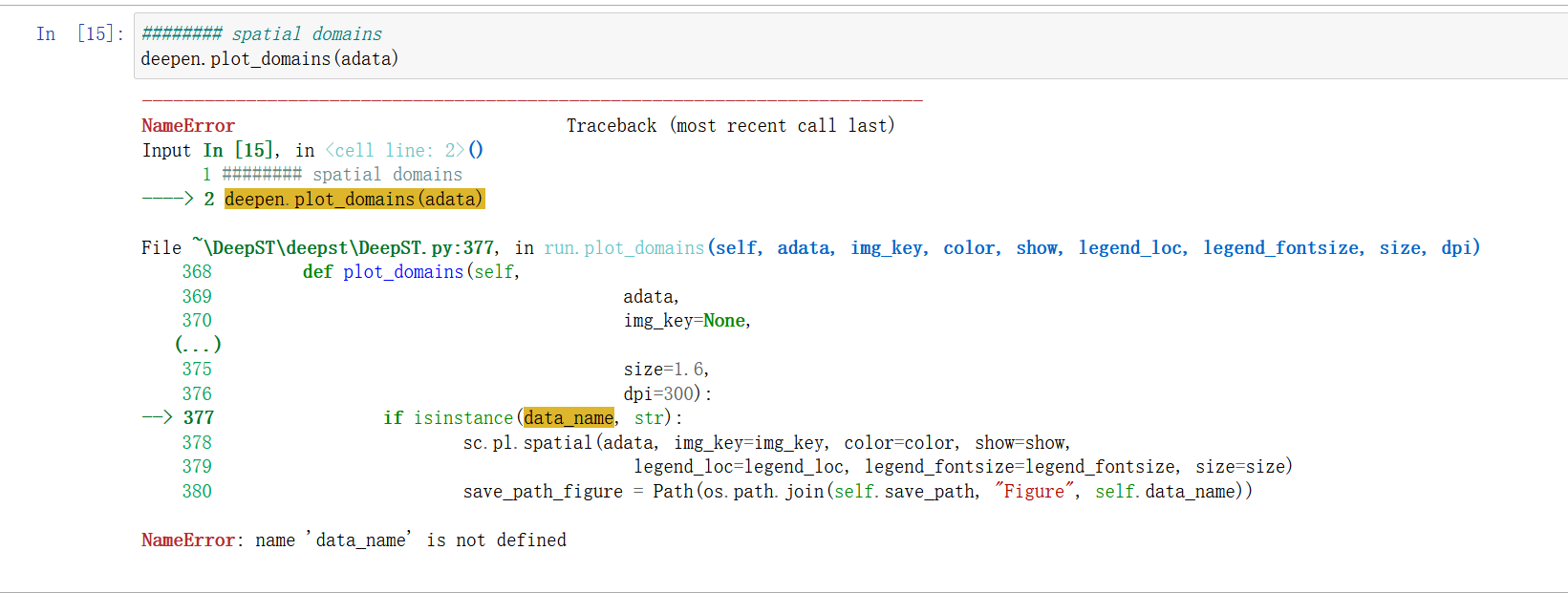
Error When I Ran The Code Deepen Plot Domains Adata Issue 1 Deepst first uses h&e staining to extract tissue morphology information through a pre trained deep learning model, and normalizes each spot’s gene expression according to the similarity of adjacent spots.
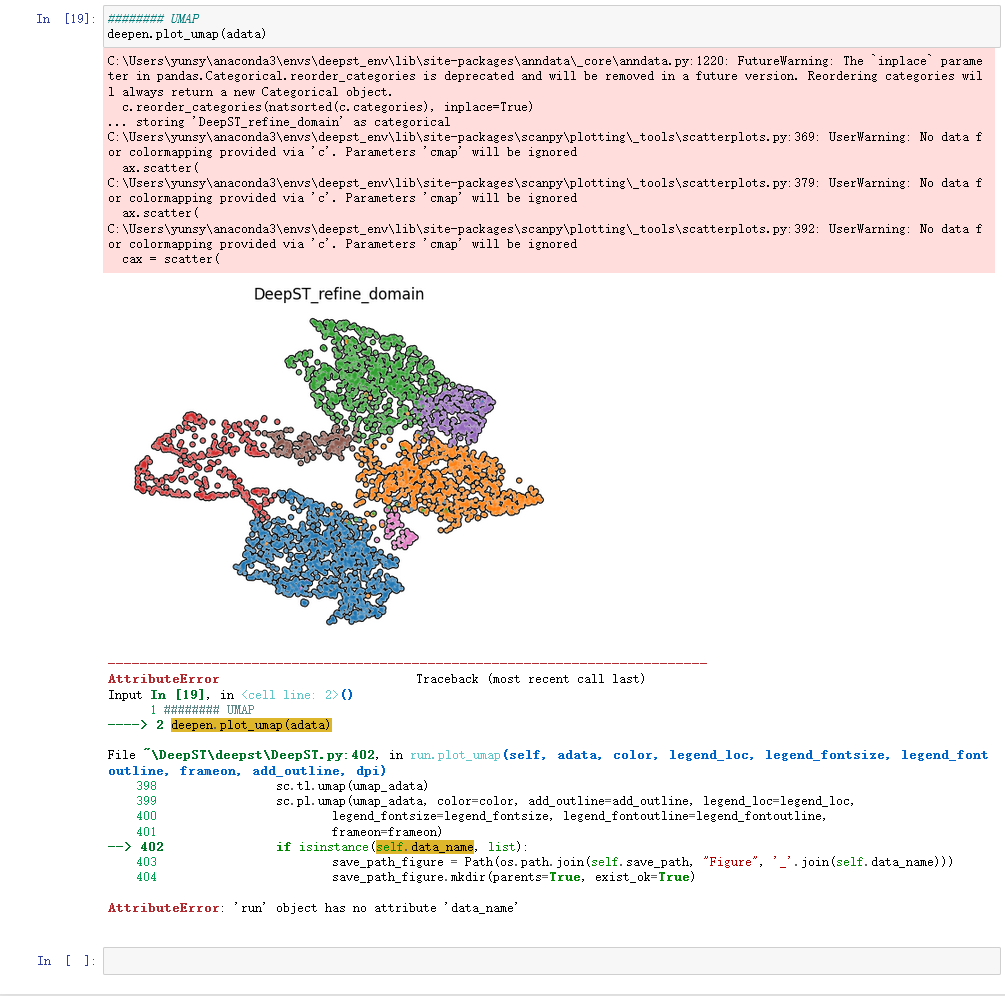
Error When I Ran The Code Deepen Plot Domains Adata Issue 1
Comments are closed.