Fish Quant

Quant Fish Quantitative Analysis Data Science Machine Learning For a detailed description of fish quant, we invited you to read our paper. the python core analysis package big fish allows you to custom build your own analysis pipeline. We provide several help files in the documents folder for new users to getting used to fish quant. we recommend working with the example data and using either step by step tutorial to familiarize yourself with the basic functionality.
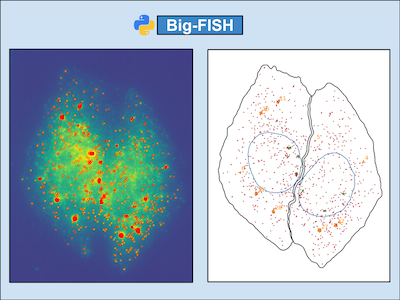
Fish Quant Getting started. read this documentation. install the imjoy plugin engine (we explain what this is just a bit further down). install the fish quant plugins. try to analyze the provided test data either with our interactive documentation (see below) or a local installation. Our user friendly package allows the user to segment nuclei and cells, detect isolated rnas, decompose dense rna clusters, quantify rna localization patterns and visualize these results both at the single cell level and variations within the cell population. Fish quant offers two solutions: (i) comparison of the integrated intensity of the transcription site to that of mature mrna and (ii) reconstruction of the transcription site signal by. A free matlab based toolbox to detect and count individual mrna transcripts in single cells by automatic analysis of 3d rna fish images. software is available freely together with extensive instructions at : code.google p fish quant .
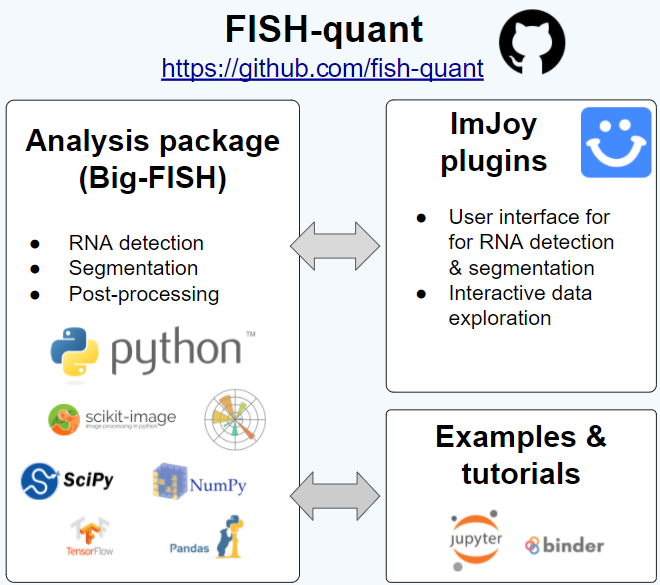
Fish Quant Fish quant offers two solutions: (i) comparison of the integrated intensity of the transcription site to that of mature mrna and (ii) reconstruction of the transcription site signal by. A free matlab based toolbox to detect and count individual mrna transcripts in single cells by automatic analysis of 3d rna fish images. software is available freely together with extensive instructions at : code.google p fish quant . Here, we introduce a python based version of our widely adopted software package fish quant (mueller et al. 2013) for the analysis of smfish images. contrary to the first version of fish quant in matlab, we address and improve on each of the specifications mentioned above. Our programs are designed to support aspiring quants from diverse backgrounds, strengths, and interests, inspiring them to confidently pursue a career in quant finance. Our user friendly package allows the user to segment nuclei and cells, detect isolated rnas, decompose dense rna clusters, quantify rna localization patterns and visualize these results both at. Fish quant matlab toolbox to analyze single molecule mrna fish data. allows counting the number of mature and nascent transcripts in 3d images. in the new version, we provide the possibility to.
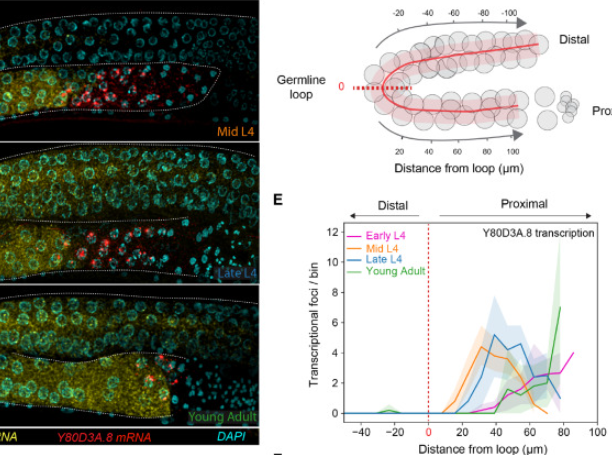
Fish Quant Here, we introduce a python based version of our widely adopted software package fish quant (mueller et al. 2013) for the analysis of smfish images. contrary to the first version of fish quant in matlab, we address and improve on each of the specifications mentioned above. Our programs are designed to support aspiring quants from diverse backgrounds, strengths, and interests, inspiring them to confidently pursue a career in quant finance. Our user friendly package allows the user to segment nuclei and cells, detect isolated rnas, decompose dense rna clusters, quantify rna localization patterns and visualize these results both at. Fish quant matlab toolbox to analyze single molecule mrna fish data. allows counting the number of mature and nascent transcripts in 3d images. in the new version, we provide the possibility to.
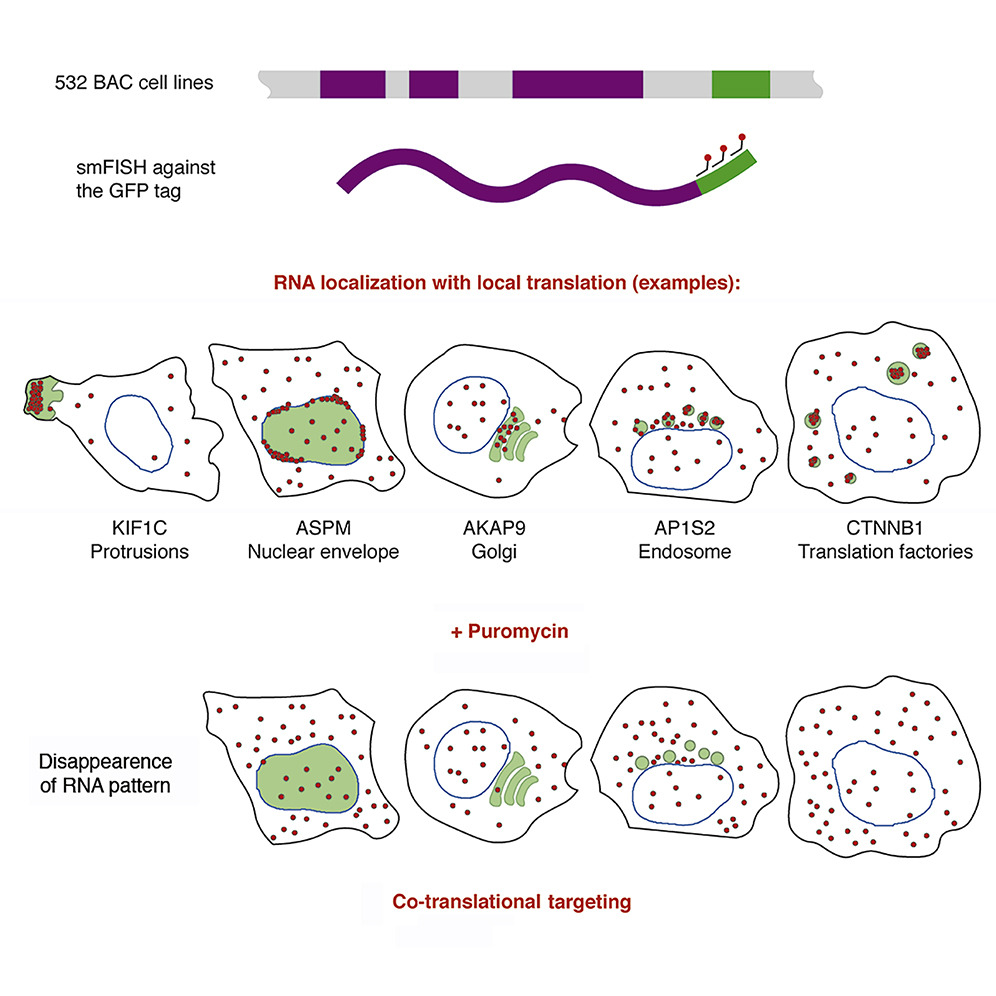
Fish Quant Our user friendly package allows the user to segment nuclei and cells, detect isolated rnas, decompose dense rna clusters, quantify rna localization patterns and visualize these results both at. Fish quant matlab toolbox to analyze single molecule mrna fish data. allows counting the number of mature and nascent transcripts in 3d images. in the new version, we provide the possibility to.
Comments are closed.