Fairifying Your Multi Omics Data

The Methods For Multi Omics Data Integration Here Simply Shows The Here, we outline a practical approach to multi modal data management and fair sharing, which are in line with the latest us and eu funders’ data sharing policies. Here, we comprehensively review state of the art multi omics integration methods with a focus on deep generative models, particularly variational autoencoders (vaes) that have been widely used for data imputation, augmentation, and batch effect correction.

The Methods For Multi Omics Data Integration Here Simply Shows The Although each omics data has individually contributed medical advances, it is their integration that may offer a more comprehensive, holistic understanding of human biology and diseases. Explore multi omics data: integration, tools, and real world impact. learn how it transforms biology and medicine. Here we develop and characterize suites of publicly available multi omics reference materials of matched dna, rna, protein and metabolites derived from immortalized cell lines from a family. We will cover the different types of multi omics integration and the options available for bulk, single cell and spatial datasets.
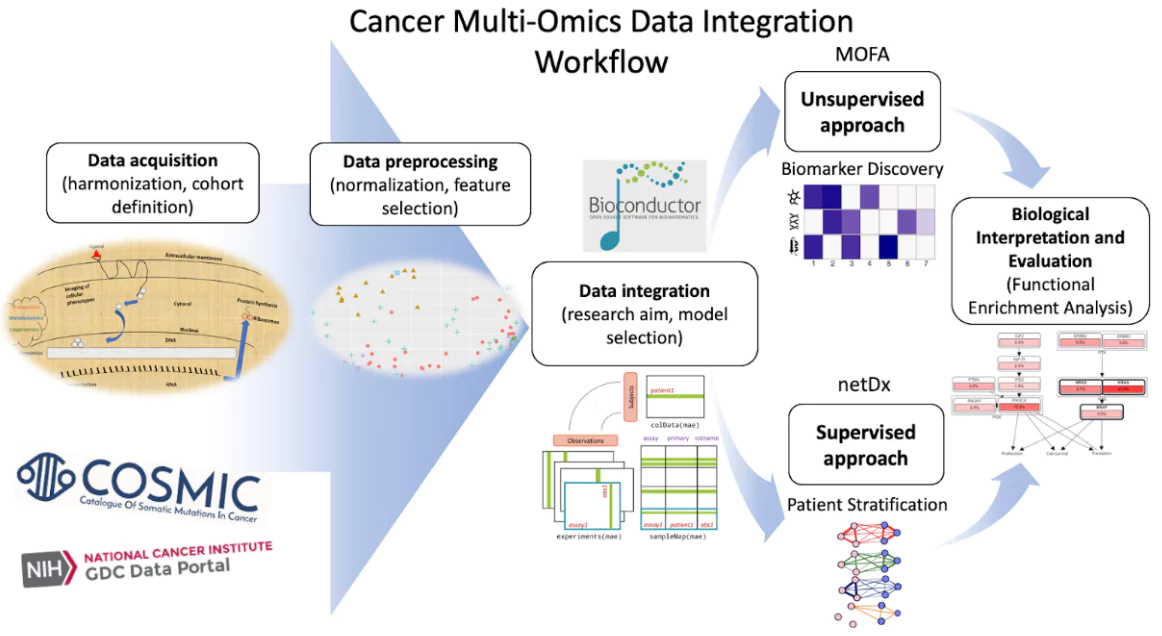
First Comprehensive Cancer Multi Omics Data Portal Pwonlyias Here we develop and characterize suites of publicly available multi omics reference materials of matched dna, rna, protein and metabolites derived from immortalized cell lines from a family. We will cover the different types of multi omics integration and the options available for bulk, single cell and spatial datasets. In this review, we categorize recent deep learning based approaches by their basic architectures and discuss their unique capabilities in relation to one another. we also discuss some emerging themes advancing the field of multi omics integration. Luckily, a complete solution to this issue exists. we will inform you on how an intuitive, coding free solution can help biologists, bioinformaticians, or translational researchers move from multi omics data to robust, reproducible insights with confidence. In this review, we collected the tools and methods that adopt integrative approach to analyze multiple omics data and summarized their ability to address applications such as disease subtyping, biomarker prediction, and deriving insights into the data. Here, we comprehensively review state of the art multi omics data integration methods with a focus on deep generative models, particularly variational autoencoders (vaes) that have been widely used for data imputation and augmentation, joint embedding creation, and batch effect correction.

Description Of Patient Matched Multi Omics Data Multi Omics Data Are In this review, we categorize recent deep learning based approaches by their basic architectures and discuss their unique capabilities in relation to one another. we also discuss some emerging themes advancing the field of multi omics integration. Luckily, a complete solution to this issue exists. we will inform you on how an intuitive, coding free solution can help biologists, bioinformaticians, or translational researchers move from multi omics data to robust, reproducible insights with confidence. In this review, we collected the tools and methods that adopt integrative approach to analyze multiple omics data and summarized their ability to address applications such as disease subtyping, biomarker prediction, and deriving insights into the data. Here, we comprehensively review state of the art multi omics data integration methods with a focus on deep generative models, particularly variational autoencoders (vaes) that have been widely used for data imputation and augmentation, joint embedding creation, and batch effect correction.
Comments are closed.