Expressed Sequence Tags

Expressed Sequence Tags Pdf Complementary Dna Gene An expressed sequence tag (est) is a short sub sequence of a cdna sequence that may be used to identify gene transcripts. learn about the history, sources, and applications of ests in genetics and genomics. Expressed sequence tags (ests) ests are short (200–500 nucleotides) dna sequences that can be used to identify a gene that is being expressed in a cell at a particular time.
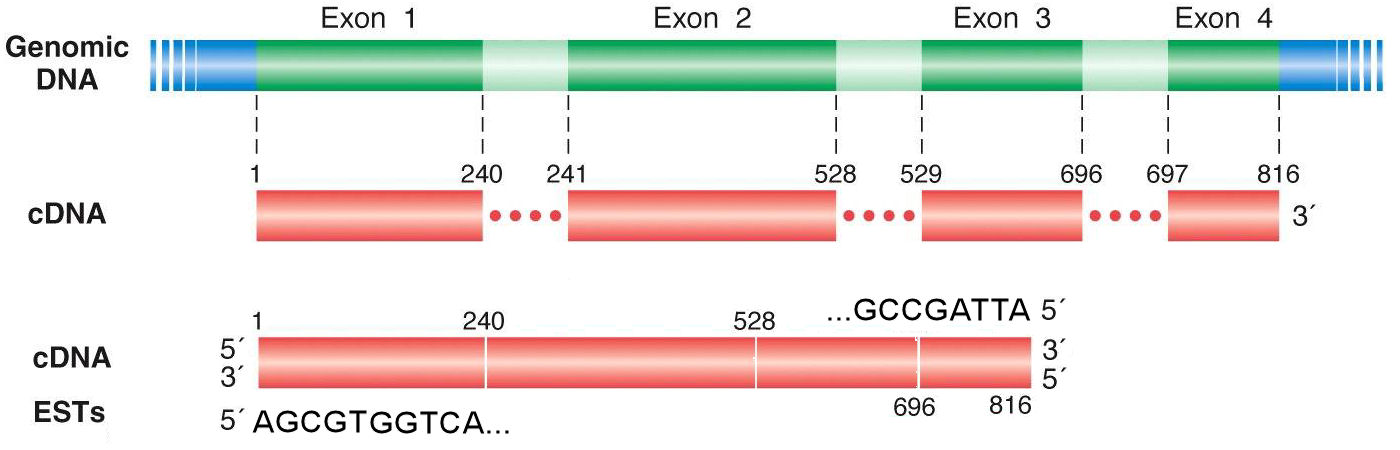
Expressed Sequence Tags An expressed sequence tag (est) refers to short sequences or partial cdnas that serve as markers for genes when the gene name is known. these sequences are named and symbolized using their sequence database accession number. Learn about expressed sequence tags (ests), their generation, applications and role in genome analysis. ests are short sequences of cdna derived from mrna that can be used to identify and map genes and genomes of different species. Ags arthur gruber 1. introduction expressed sequence tags (ests) are short sequence reads, typically within the range of 00–700 bp (see fig. 1), obtained f. om randomly selected cdna clones. the concept was first introduced as a cost effective approach for the rapid discovery and charac. Later, these cdna clones are read in a single pass from either end, resulting in a subsequence of the original clone called an expressed sequence tag or simply est.

Ppt Expressed Sequence Tags Powerpoint Presentation Free Download Ags arthur gruber 1. introduction expressed sequence tags (ests) are short sequence reads, typically within the range of 00–700 bp (see fig. 1), obtained f. om randomly selected cdna clones. the concept was first introduced as a cost effective approach for the rapid discovery and charac. Later, these cdna clones are read in a single pass from either end, resulting in a subsequence of the original clone called an expressed sequence tag or simply est. Expressed sequence tags (ests) are fragments of mrna sequences derived through single sequencing reactions performed on randomly selected clones from cdna libraries. to date, over 45 million ests have been generated from over 1400 different species of eukaryotes. The analysis of expressed sequence tag (est) datasets offers a rapid and cost effective approach to elucidate the transcriptome of an organism, but requiring several computational methods for assembly and annotation. Sequencing a short segment (<100 bases) at one or both ends of each cdna produces a "tag" with enough information to identify the expressed gene. these expressed sequence tags (ests) from a novel organism or tissue type can be compared with sequences from a well characterized genome. Expressed sequence tags (ests) are short subsequences of genes, typically 200 to 500 base pairs in length, that are derived from cloned dna (cdna) and used to identify gene expression.

Figure 1 From Lecture 33 Expressed Sequence Tags Semantic Scholar Expressed sequence tags (ests) are fragments of mrna sequences derived through single sequencing reactions performed on randomly selected clones from cdna libraries. to date, over 45 million ests have been generated from over 1400 different species of eukaryotes. The analysis of expressed sequence tag (est) datasets offers a rapid and cost effective approach to elucidate the transcriptome of an organism, but requiring several computational methods for assembly and annotation. Sequencing a short segment (<100 bases) at one or both ends of each cdna produces a "tag" with enough information to identify the expressed gene. these expressed sequence tags (ests) from a novel organism or tissue type can be compared with sequences from a well characterized genome. Expressed sequence tags (ests) are short subsequences of genes, typically 200 to 500 base pairs in length, that are derived from cloned dna (cdna) and used to identify gene expression.

What Is The Difference Between Expressed Sequence Tags And Sequence Sequencing a short segment (<100 bases) at one or both ends of each cdna produces a "tag" with enough information to identify the expressed gene. these expressed sequence tags (ests) from a novel organism or tissue type can be compared with sequences from a well characterized genome. Expressed sequence tags (ests) are short subsequences of genes, typically 200 to 500 base pairs in length, that are derived from cloned dna (cdna) and used to identify gene expression.
Comments are closed.