Efficient Sequence Alignment With Dynamic Programming Peerdh

Lecture 7 Dynamic Programming Global Sequence Alignment Pdf In this article, we will look at how to implement dynamic programming for sequence alignment in python, focusing on the needleman wunsch algorithm for global alignment and the smith waterman algorithm for local alignment. Efficient implementation of algorithms for optimal resolution of the peerwise sequence alignment problem. sequence alignment has a great biological interest being a key to finding important regions in the genome, determining its functions and uncovering evolutionary forces.
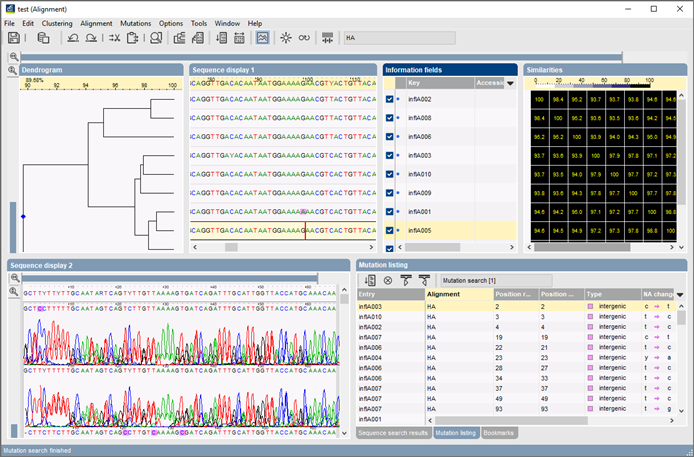
Efficient Sequence Alignment With Dynamic Programming Peerdh An alignment is an assignment of gaps to positions 0, , m in x, and 0, , n in y, so as to line up each letter in one sequence with either a letter, or a gap in the other sequence. Giving two sequences seq1 and seq2 instead of determining the similarity between sequences as a whole, dynamic programming allows construction of the solution by determining all parallels between random prefixes of the two sequences. We compare 2 pairwise methods (dynamic programming and local search) and implement a multiple alignment task based on them. then we give a demo of showing how to query the whole genome of shewanella, a marine bacterium whose o linked glycosylation pathway is not known, for protein similar to pseb. Given two groups a and b of aligned sequences, this algorithm uses dynamic programming and the sum of pairs objective function to determine an optimal alignment c of a and b.

Efficient Data Structures For Sequence Alignment Peerdh We compare 2 pairwise methods (dynamic programming and local search) and implement a multiple alignment task based on them. then we give a demo of showing how to query the whole genome of shewanella, a marine bacterium whose o linked glycosylation pathway is not known, for protein similar to pseb. Given two groups a and b of aligned sequences, this algorithm uses dynamic programming and the sum of pairs objective function to determine an optimal alignment c of a and b. To this end, we introduce alignment in memory (aim), a framework for pim based sequence alignment that targets the upmem system. aim dispatches a large number of sequence pairs across diferent memory modules and aligns each pair using compute cores within the memory module where the pair resides. But there is no method that can solve the problem of multiple sequence alignment efficiently and optimally.
dynamic programming method has been proven to handle pairwise sequence alignment effective and efficiently at both global and local alignment. This is where dynamic programming comes into play. it provides an efficient way to align sequences by breaking the problem into smaller, manageable subproblems. A scheme for parallel implementation of the dynamic programming multiple sequence alignment is presented, based on a peer to peer design and a multidimensional array indexing method.

An Introduction To Dna Sequence Alignment Algorithms Exploring Dynamic To this end, we introduce alignment in memory (aim), a framework for pim based sequence alignment that targets the upmem system. aim dispatches a large number of sequence pairs across diferent memory modules and aligns each pair using compute cores within the memory module where the pair resides. But there is no method that can solve the problem of multiple sequence alignment efficiently and optimally.
dynamic programming method has been proven to handle pairwise sequence alignment effective and efficiently at both global and local alignment. This is where dynamic programming comes into play. it provides an efficient way to align sequences by breaking the problem into smaller, manageable subproblems. A scheme for parallel implementation of the dynamic programming multiple sequence alignment is presented, based on a peer to peer design and a multidimensional array indexing method.

Scoring Algorithms In Sequence Alignment Peerdh This is where dynamic programming comes into play. it provides an efficient way to align sequences by breaking the problem into smaller, manageable subproblems. A scheme for parallel implementation of the dynamic programming multiple sequence alignment is presented, based on a peer to peer design and a multidimensional array indexing method.
Comments are closed.