Developing Reproducible Workflows Collaboratively
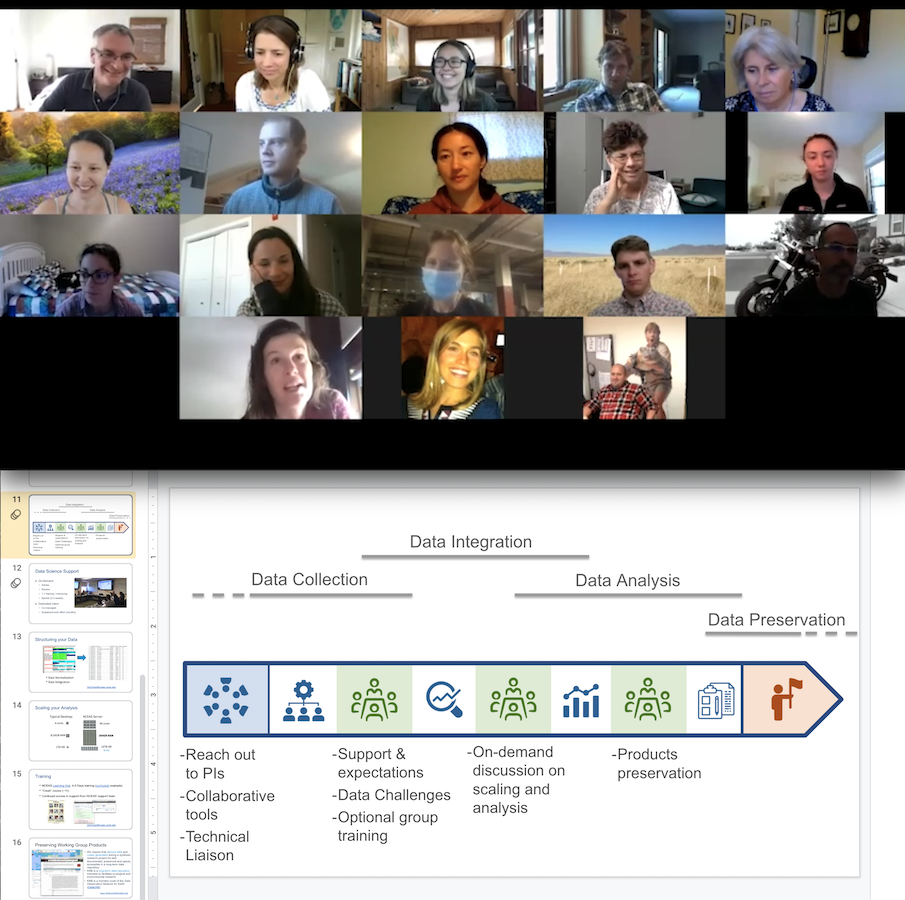
Developing Reproducible Workflows Collaboratively In this module, we wanted to empower multi institutional and cross sectoral science teams to collaboratively develop reproducible workflows for data driven projects. In this module, we wanted to empower multi institutional and cross sectoral science teams to collaboratively develop reproducible workflows for data driven projects.

Developing Reproducible Workflows Collaboratively In this module, we wanted to empower multi institutional and cross sectoral science teams to collaboratively develop reproducible workflows for data driven projects. This manuscript shares the lessons learned from providing scientific computing support to over 600 researchers and discipline experts, helping them develop reproducible and scalable analytical workflows to process large amounts of heterogeneous data. Automation of computational workflows is essential for reproducibility in robotic experimental systems. a task choreographer based on directed acyclic graphs favors collaborative parallel self driving experimentation. containerized applications facilitate integration and co experimentation. Nextflow | nextflow is a leading open source workflow orchestrator that simplifies writing and deploying data intensive pipelines at scale on any infrastructure. with nextflow, scientists and bioinformaticians can easily create and share reliable, reproducible scientific workflows or adapt existing workflows to nextflow using their preferred scripting language. unlike other tools where.
Developing Reproducible Workflows Collaboratively Automation of computational workflows is essential for reproducibility in robotic experimental systems. a task choreographer based on directed acyclic graphs favors collaborative parallel self driving experimentation. containerized applications facilitate integration and co experimentation. Nextflow | nextflow is a leading open source workflow orchestrator that simplifies writing and deploying data intensive pipelines at scale on any infrastructure. with nextflow, scientists and bioinformaticians can easily create and share reliable, reproducible scientific workflows or adapt existing workflows to nextflow using their preferred scripting language. unlike other tools where. We will cover a number of processes, systems, and practices that you should adopt to help ensure that your work is efficient, your processing steps traceable and repeatable, your analyses and findings reproducible, and your data and processing scripts amenable to sharing and open science. What has been the response from scientists using agilent automation for drug discovery? agilent automation is used across the adc discovery and development process to enable researchers to scale throughput, remove variability from complex workflows and generate more reproducible data with less hands on effort. We show how to create a scalable, reusable, and shareable workflow using four different workflow engines: the common workflow language (cwl), guix workflow language (gwl), snakemake, and nextflow. each of which can be run in parallel. How i use nix and direnv to create isolated, reproducible development environments. no more "works on my machine" — just clean, version pinned shells that activate automatically. tagged with nixos, development, workflow.
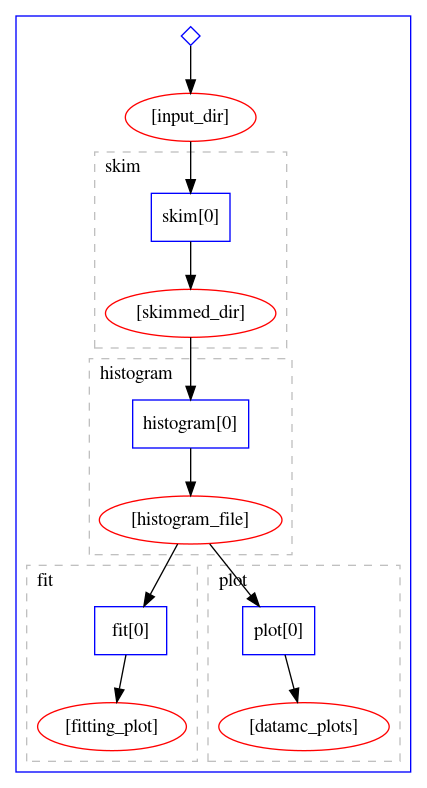
Developing Parallel Workflows Reproducible Analyses We will cover a number of processes, systems, and practices that you should adopt to help ensure that your work is efficient, your processing steps traceable and repeatable, your analyses and findings reproducible, and your data and processing scripts amenable to sharing and open science. What has been the response from scientists using agilent automation for drug discovery? agilent automation is used across the adc discovery and development process to enable researchers to scale throughput, remove variability from complex workflows and generate more reproducible data with less hands on effort. We show how to create a scalable, reusable, and shareable workflow using four different workflow engines: the common workflow language (cwl), guix workflow language (gwl), snakemake, and nextflow. each of which can be run in parallel. How i use nix and direnv to create isolated, reproducible development environments. no more "works on my machine" — just clean, version pinned shells that activate automatically. tagged with nixos, development, workflow.
Comments are closed.