Deep Learning To Integrate Histology With Spatial Transcriptomics

Deep Learning To Integrate Histology With Spatial Transcriptomics In this study, we identify 22 dl methods for spatial transcriptomics integration with other modalities, including histology images, scrna seq, chromatin images, and other spatial omics data. Gist provides a generalizable framework for integrating histology with spatial transcriptome analysis, revealing novel insights into spatial organization and functional dynamics.

Pdf Generalization Of Deep Learning Models For Predicting Spatial In this study, we systematically review the applications of dl in integrating spatial transcriptomics data with other modalities. Here, we propose staig, a deep learning model that integrates gene expression, spatial coordinates, and histological images using graph contrastive learning coupled with high performance. Abstract spatial transcriptomics (st) is a powerful assay to capture gene expression in tissue context. however, due to the limitation of resolution, most existing st datasets remain at multicellular resolution which hinders comprehensive understanding of. To address this, we develop a multi scale convolutional deep learning framework, hist, which utilizes st to learn the relationship between spatially resolved gene expression profiles (geps) and histological morphology.

Spatial Pathomics Toolkit For Quantitative Analysis Of Podocyte Nuclei Abstract spatial transcriptomics (st) is a powerful assay to capture gene expression in tissue context. however, due to the limitation of resolution, most existing st datasets remain at multicellular resolution which hinders comprehensive understanding of. To address this, we develop a multi scale convolutional deep learning framework, hist, which utilizes st to learn the relationship between spatially resolved gene expression profiles (geps) and histological morphology. These methods integrate spatial multi modal data using a deep learning model to derive spatially embedded features that capture underlying biological functions and spatial tissue patterns. To address this challenge, we propose spacrd, a transfer learning based method that deeply integrates histology images and st data to enable reliable ctr detection across diverse samples, platforms, and batches. We introduce deepspot, a novel deep learning model that predicts spatial transcriptomics from h&e images. deepspot employs a deep set neural network to model spots as bags of sub spots and integrates multi level tissue details and spatial context. I will demonstrate st net on breast cancer data, and show how it characterizes tumor spatial heterogeneity and quantifies tumor immune interactions. i will then discuss how similar algorithms can capture complex morphological changes over time of individual cells.

Deep Learning Framework Advances Tissue Analysis In Spatial Transcriptomics These methods integrate spatial multi modal data using a deep learning model to derive spatially embedded features that capture underlying biological functions and spatial tissue patterns. To address this challenge, we propose spacrd, a transfer learning based method that deeply integrates histology images and st data to enable reliable ctr detection across diverse samples, platforms, and batches. We introduce deepspot, a novel deep learning model that predicts spatial transcriptomics from h&e images. deepspot employs a deep set neural network to model spots as bags of sub spots and integrates multi level tissue details and spatial context. I will demonstrate st net on breast cancer data, and show how it characterizes tumor spatial heterogeneity and quantifies tumor immune interactions. i will then discuss how similar algorithms can capture complex morphological changes over time of individual cells.

Hest 1k A Dataset For Spatial Transcriptomics And Histology Image We introduce deepspot, a novel deep learning model that predicts spatial transcriptomics from h&e images. deepspot employs a deep set neural network to model spots as bags of sub spots and integrates multi level tissue details and spatial context. I will demonstrate st net on breast cancer data, and show how it characterizes tumor spatial heterogeneity and quantifies tumor immune interactions. i will then discuss how similar algorithms can capture complex morphological changes over time of individual cells.
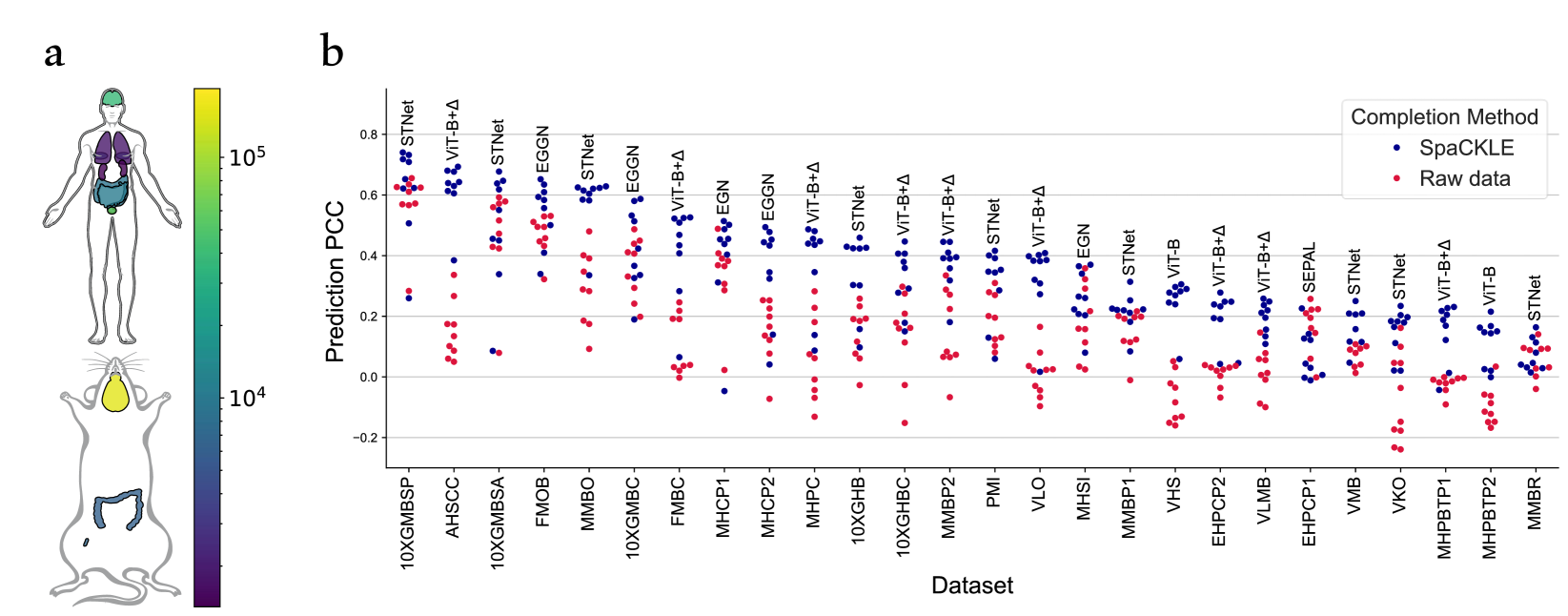
High Resolution Spatial Transcriptomics From Histology Images Using
Comments are closed.