Data Types Used For Snp Data Download Table

Data Types Used For Snp Data Download Table We apply clumping and thresholding to obtain approximately independent snp lists using ld patterns from topmed. you can browse a trait, retrieve related traits, and download snp lists ready for local prs computation. These analyses are based on sets of genome coordinates representing transcriptionally active gene loci generated for example by micro array or rna seq experiments and sets of coordinates of.

Data Types Used For Snp Data Download Table Snp tables in odi 12c design overview this document contains a list of over 100 table names prefixed with "snp " that are related to oracle data integrator (odi) metadata stored in repositories. We recommend downloading igsr data via globus (instructions). data can also be downloaded via ftp (ftp site) or aspera (instructions). ftp access is subject to rate limiting, so it is essential to download local copies for your work rather than try to access via range queries remotely. It includes descriptions of over 50 different tables that define items like transformations, mappings, projects, and other metadata stored in the repository. the tables use a common prefix of "snp " and columns are defined with data types. Download a full copy of the gwas catalog in spreadsheet format as well as current and older versions of the gwas diagram in svg format. documentation and access to full summary statistics for gwas catalog studies where available. submit summary statistics to gwas catalog.

Snp Data Analysis Allgenetics It includes descriptions of over 50 different tables that define items like transformations, mappings, projects, and other metadata stored in the repository. the tables use a common prefix of "snp " and columns are defined with data types. Download a full copy of the gwas catalog in spreadsheet format as well as current and older versions of the gwas diagram in svg format. documentation and access to full summary statistics for gwas catalog studies where available. submit summary statistics to gwas catalog. In animal snpatlas, users can search the functional annotation of snps, perform online genotype imputation, explore and visualize ld information, browse variant information using the genome browser and download snp datasets for each species. Prs csx is a python based command line tool that integrates gwas summary statistics and external ld reference panels from multiple populations to improve cross population polygenic prediction. posterior snp effect sizes are inferred under coupled continuous shrinkage (cs) priors across populations. We describe updates to the rice snp seek database since its first release. we ran a new snp calling pipeline followed by filtering that resulted in complete, base, filtered and core snp datasets. Whole genome collections of snp and short indel variants are available for 52 inbred strains. the variants are derived from illumina short read sequencing data generated between 2013 2021 from a mixture of read lengths (100 150bp).

Snp Data Pdf In animal snpatlas, users can search the functional annotation of snps, perform online genotype imputation, explore and visualize ld information, browse variant information using the genome browser and download snp datasets for each species. Prs csx is a python based command line tool that integrates gwas summary statistics and external ld reference panels from multiple populations to improve cross population polygenic prediction. posterior snp effect sizes are inferred under coupled continuous shrinkage (cs) priors across populations. We describe updates to the rice snp seek database since its first release. we ran a new snp calling pipeline followed by filtering that resulted in complete, base, filtered and core snp datasets. Whole genome collections of snp and short indel variants are available for 52 inbred strains. the variants are derived from illumina short read sequencing data generated between 2013 2021 from a mixture of read lengths (100 150bp).
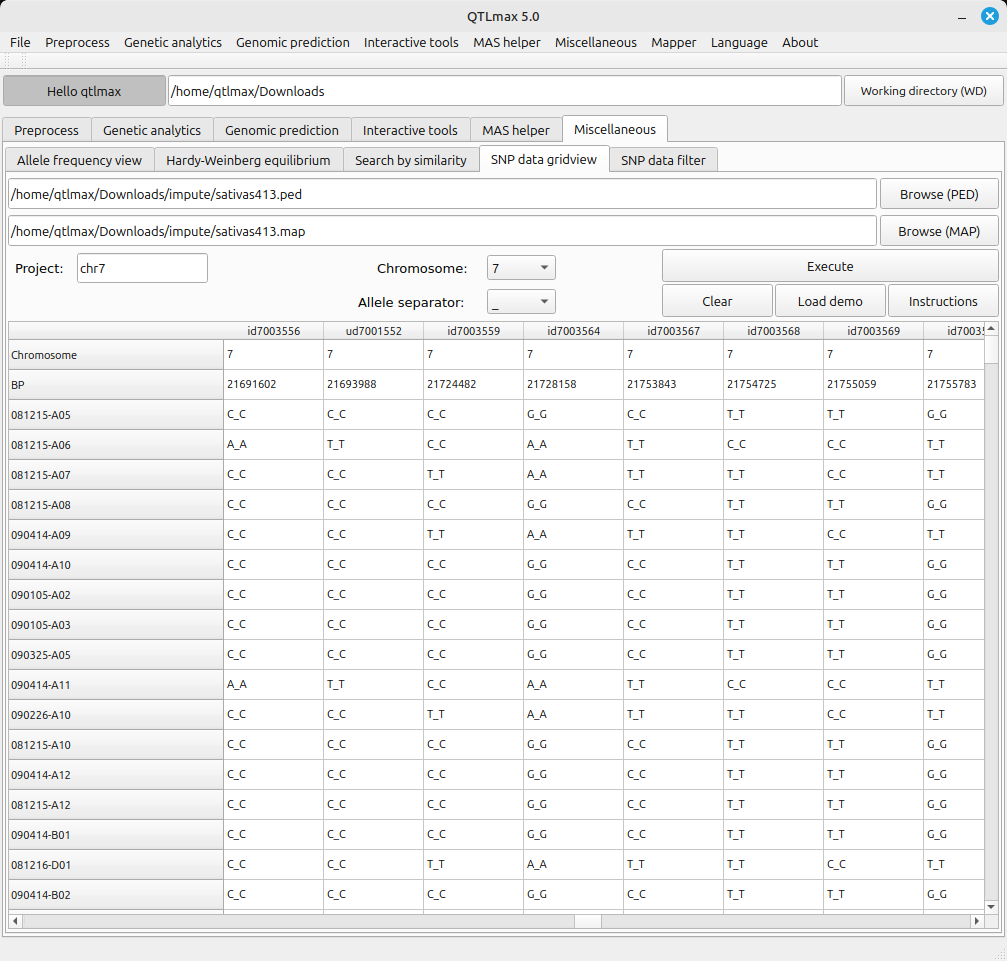
How To Display Snp Data As A Table Qtlmax Open We describe updates to the rice snp seek database since its first release. we ran a new snp calling pipeline followed by filtering that resulted in complete, base, filtered and core snp datasets. Whole genome collections of snp and short indel variants are available for 52 inbred strains. the variants are derived from illumina short read sequencing data generated between 2013 2021 from a mixture of read lengths (100 150bp).
Comments are closed.