Converting Microarray Data To Vcf Format In Varseq Pgx

Converting Microarray Data To Vcf Format In Varseq Pgx Golden helix has developed a process of converting the array data to the vcf structure necessary to build a varseq project. the landscape of array data formats may vary across platforms, but the strategy for conversion remains constant. Our latest blog dives into the "how" and "why" behind this transformation, showcasing the versatility of varseq in handling diverse genetic data types.

Converting Microarray Data To Vcf Format In Varseq Pgx Did you know that converting your microarray data to vcf format can open up new possibilities for pharmacogenomics (pgx) analysis? our latest blog dives into the "how" and "why" behind this transformation, showcasing the versatility of varseq in handling diverse genetic data types. A set of tools to convert illumina and affymetrix dna microarray intensity data files into vcf files without using microsoft windows. you can use the final output to run the pipeline to detect mosaic chromosomal alterations. Hello everyone, i have only one microarray data which contains one .cel file and another intensity .csv file. i'm searching for converting any of the two files into vcf format. Variant calling format is a tab delimited text file that is used to describe single nucleotide variants (snvs) as well as insertions, deletions, and other sequence variations.
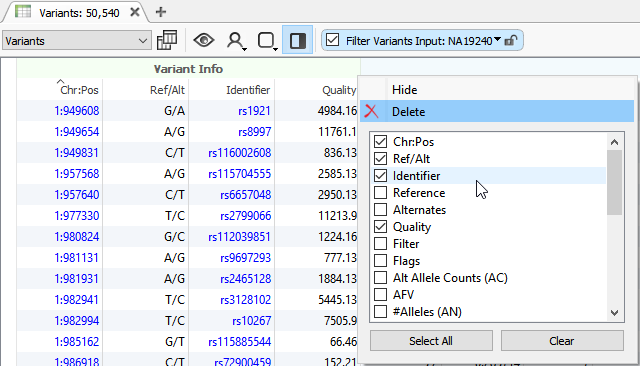
How To Varseq Vcf Import The Golden Helix Blog Hello everyone, i have only one microarray data which contains one .cel file and another intensity .csv file. i'm searching for converting any of the two files into vcf format. Variant calling format is a tab delimited text file that is used to describe single nucleotide variants (snvs) as well as insertions, deletions, and other sequence variations. The information provided by the gatk tools that infer variation from high throughput sequencing data, such as the haplotypecaller, is especially complex. this document describes the key features and annotations that you need to know about in order to understand vcf files output by the gatk tools. For this tutorial, we will use bcftools which is designed by the same team behind samtools they are part of the same pipeline. bcftools is itself a comprehensive pipeline and produces a variant call format (vcf) that is used in many downstream analyses. Acli also uses a command line interface to generate gtc and vcf files from idats. it is recommended to use dragen array which has additional features compared to acli. The aim is to get some hands on experience working with vcf files, practice some of the considerations in producing a subset of high quality snps, and to explore the data structure using a principal component analysis (pca). the aim is also to apply and expand on the unix and r skills that you learned in the previous activities: unix and r.

How To Varseq Vcf Import The Golden Helix Blog The information provided by the gatk tools that infer variation from high throughput sequencing data, such as the haplotypecaller, is especially complex. this document describes the key features and annotations that you need to know about in order to understand vcf files output by the gatk tools. For this tutorial, we will use bcftools which is designed by the same team behind samtools they are part of the same pipeline. bcftools is itself a comprehensive pipeline and produces a variant call format (vcf) that is used in many downstream analyses. Acli also uses a command line interface to generate gtc and vcf files from idats. it is recommended to use dragen array which has additional features compared to acli. The aim is to get some hands on experience working with vcf files, practice some of the considerations in producing a subset of high quality snps, and to explore the data structure using a principal component analysis (pca). the aim is also to apply and expand on the unix and r skills that you learned in the previous activities: unix and r.
Comments are closed.