Computational Genomics Tutorial Genomics Tutorial 2020 2 0 Documentation
Computational Genomics Assignment Point This is an introductory tutorial for learning computational genomics mostly on the linux command line. you will learn how to analyse next generation sequencing (ngs) data. This is an introductory tutorial for learning computational genomics mostly on the linux command line. you will learn how to analyse next generation sequencing (ngs) data.
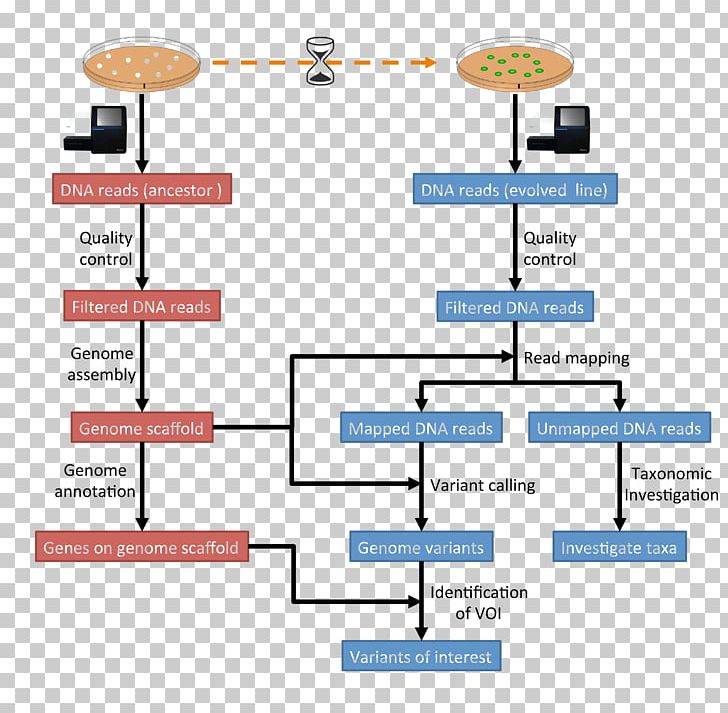
Computational Genomics Computational Biology Research Bioinformatics This is an introductory tutorial for learning computational genomics mostly on the linux command line. you will learn how to analyse next generation sequencing (ngs) data. This is an introductory tutorial for learning genomics mostly on the linux command line. should you need to refresh your knowledge about either linux or the command line, have a look here. Sphinx documentation source for a computational genomics tutorial. For this tutorial we will assume little background knowledge, except for a basic familiarity with the linux operating system and the cloud. we will cover the basics of how genomic dna libraries are generated and sequenced, and the principles behind short read paired end sequencing.
Computational Structural Genomics Github Sphinx documentation source for a computational genomics tutorial. For this tutorial we will assume little background knowledge, except for a basic familiarity with the linux operating system and the cloud. we will cover the basics of how genomic dna libraries are generated and sequenced, and the principles behind short read paired end sequencing. This is an introductory tutorial for learning computational genomics mostly on the linux command line. you will learn how to analyse next generation sequencing (ngs) data. These tutorials cover the entire research workflow — from environment setup and data management to pipeline development, quality control, visualization, and collaboration. Rob, liz dinsdale, tom jeffries, bruno gomez gil, jim mitchell, and several other colleagues and friends have been teaching genomics and metagenomics for a long time. they have written this manual over the course of several years, and in a variety of formats. Free and easy to use, the open science framework supports the entire research lifecycle: planning, execution, reporting, archiving, and discovery.
Computational Genomics Tse Lab At Stanford University This is an introductory tutorial for learning computational genomics mostly on the linux command line. you will learn how to analyse next generation sequencing (ngs) data. These tutorials cover the entire research workflow — from environment setup and data management to pipeline development, quality control, visualization, and collaboration. Rob, liz dinsdale, tom jeffries, bruno gomez gil, jim mitchell, and several other colleagues and friends have been teaching genomics and metagenomics for a long time. they have written this manual over the course of several years, and in a variety of formats. Free and easy to use, the open science framework supports the entire research lifecycle: planning, execution, reporting, archiving, and discovery.
Comments are closed.