Clustering As Dimensionality Reduction

Clustering Before Or After Dimensionality Reduction By following the best practices outlined in this article, you can effectively cluster high dimensional data and derive meaningful insights from complex datasets. Dimensionality reduction (dr) simplifies complex data from genomics, imaging, sensors, and language into interpretable forms that support visualization, clustering, and modeling.
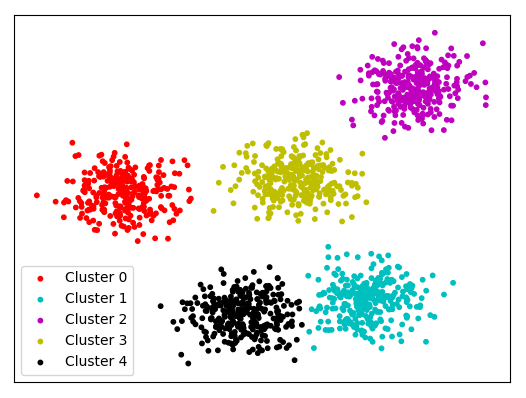
K Means Clustering On Spherical Data 1v2 This article delves into key methods beyond basic clustering and explores how dimensionality reduction simplifies high dimensional datasets while preserving critical insights. This article delves into the principles, methodologies, and practical applications of feature selection and dimensionality reduction in the context of clustering. In this work, we investigate conditions under which two types of clusters (contiguous and density clusters) are preserved after a one dimensional embedding through isomap. To address these limitations we consider a family of two stage models that implement dimensionality reduction with a linear gaussian model (lgm) such as pca or factor analysis (fa), and clustering with a mixture of gaussians (mog).

The Three Main Tasks And Their Algorithms In Unsupervised Learning In this work, we investigate conditions under which two types of clusters (contiguous and density clusters) are preserved after a one dimensional embedding through isomap. To address these limitations we consider a family of two stage models that implement dimensionality reduction with a linear gaussian model (lgm) such as pca or factor analysis (fa), and clustering with a mixture of gaussians (mog). This work proposes a new technique for reducing the number of variables in datasets. the procedure is based on combining variable cluster analysis and canonical correlation analysis to determine synthetic variables that are representative of the clusters. High dimensional data is everywhere — gene expression panels with tens of thousands of features, image embeddings with hundreds of channels, and survey instruments with dozens of scales. visualizing or clustering such data demands dimensionality reduction, but choosing the right method is not trivial. the three dominant approaches are pca, t sne, and umap. each makes fundamentally different. You will learn how hierarchical clustering builds a tree like structure (dendrogram) to group similar data points, and how dimensionality reduction techniques help reduce the number of features. In machine learning, clustering and dimensionality reduction are two of the most important methods for dealing with complex data. clustering groups data by similarities, which reveals patterns in the data, whereas dimensionality reduction reduces the number of variables, thus making the data easier to work with.
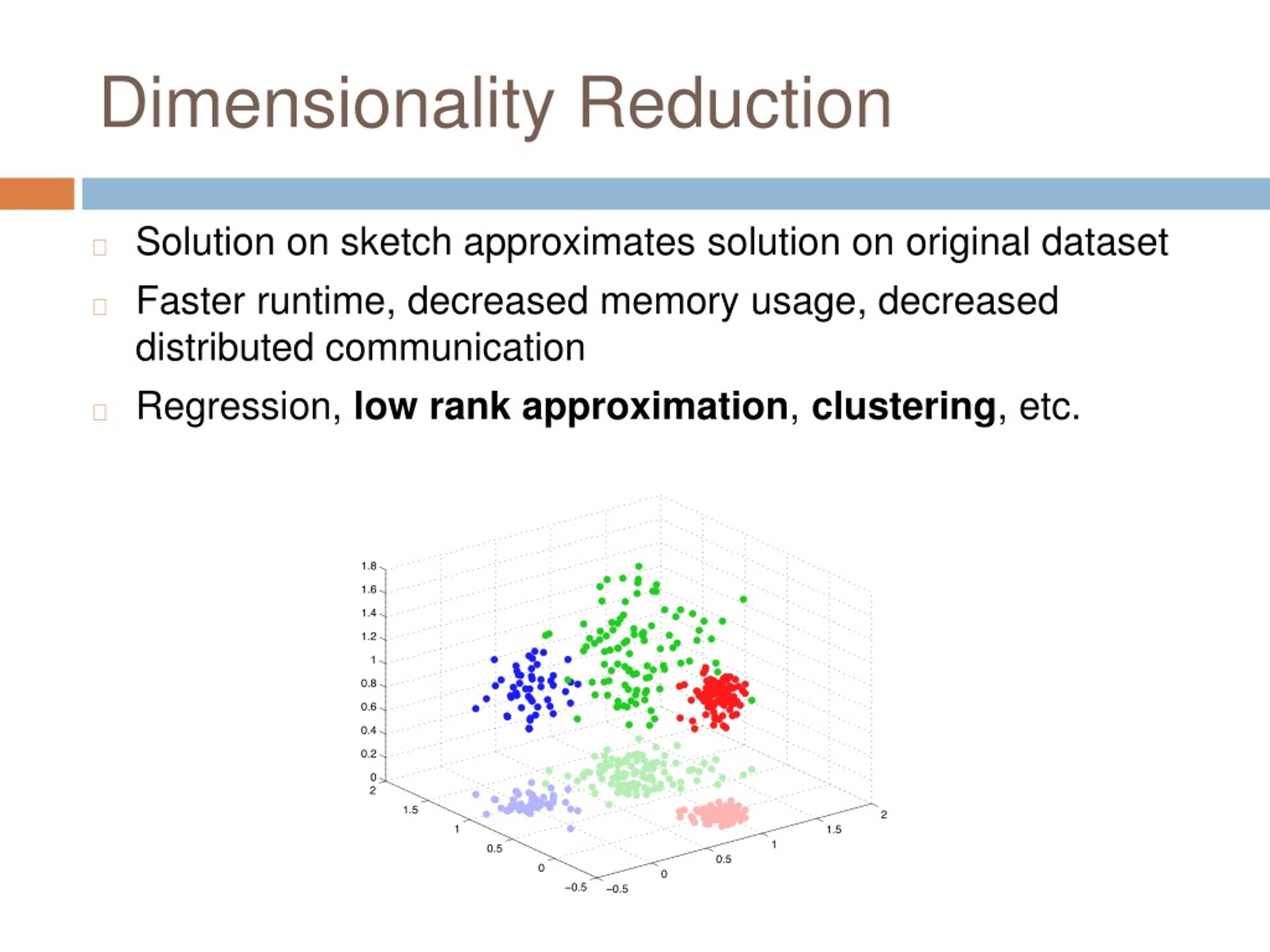
Ppt Dimensionality Reduction For K Means Clustering And Low Rank This work proposes a new technique for reducing the number of variables in datasets. the procedure is based on combining variable cluster analysis and canonical correlation analysis to determine synthetic variables that are representative of the clusters. High dimensional data is everywhere — gene expression panels with tens of thousands of features, image embeddings with hundreds of channels, and survey instruments with dozens of scales. visualizing or clustering such data demands dimensionality reduction, but choosing the right method is not trivial. the three dominant approaches are pca, t sne, and umap. each makes fundamentally different. You will learn how hierarchical clustering builds a tree like structure (dendrogram) to group similar data points, and how dimensionality reduction techniques help reduce the number of features. In machine learning, clustering and dimensionality reduction are two of the most important methods for dealing with complex data. clustering groups data by similarities, which reveals patterns in the data, whereas dimensionality reduction reduces the number of variables, thus making the data easier to work with.
Comments are closed.