Cellprofiler Output During Processing Whitelader

Cellprofiler Output During Processing Redlader You can save any of the processed images created by cellprofiler during the analysis using this module. you can choose from many different image formats for saving your files. The tutorial is a step by step guide and contains example data, cellprofiler pipelines and a machine learning script (in python) which can serve as a starting point when analyzing your own ifc data.

Cellprofiler Output During Processing Whitelader I was able to get the output data after running a smaller data set. a new issue i have is running a full 96 well data with almost 900 image sets and file processing. i get a warning from cell profiler that exporttospreadsheet isn’t a good option and its likely to fail but exporttodatabase is better. i have not been able to get that to run. The nih has published a introductory chapter of “best practices” for image based high content screening (in which cellprofiler is mentioned) as part of the assay guidance manual, and our group has published a more advanced follow up chapter on image analysis methods. A pipeline is a sequential set of image analysis measurement and processing modules. each module is how we give cellprofiler instructions (one at a time) in order to process, analyze and export any images and data. To change the default folder where data and images are output from cellprofiler, select the “cellprofiler” dropdown menu and select “preferences”. then select “browse” next to the “default output folder”.
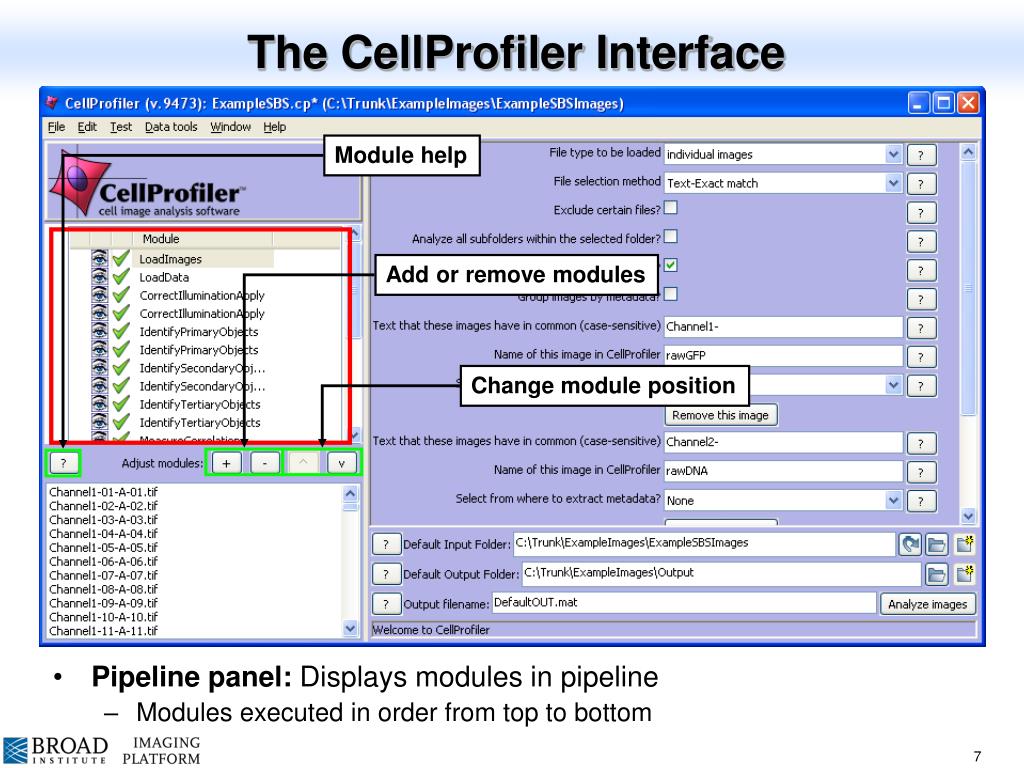
Cellprofiler Output During Processing Whitelader A pipeline is a sequential set of image analysis measurement and processing modules. each module is how we give cellprofiler instructions (one at a time) in order to process, analyze and export any images and data. To change the default folder where data and images are output from cellprofiler, select the “cellprofiler” dropdown menu and select “preferences”. then select “browse” next to the “default output folder”. Using cellprofiler and fcs express 4 image cytometry: joint webinar with denovo software on using cellprofiler to peform basic image assays and generate data compatible with export into fcs express. The steps for exporting data from cellprofiler have been outlined by illustrating how to add images, extract metadata from files, and modify the required modules according to the section1pipeline.cpproj. When specified, cellprofiler will output one line to the terminal per job to be run. this output should be further processed to generate a script that can invoke the jobs in a cluster computing context. Cellprofiler can be used to analyze the resulting images from imaging flow cytometry, whether brightfield, darkfield, or fluorescence. the ifc website page has further details on this workflow.
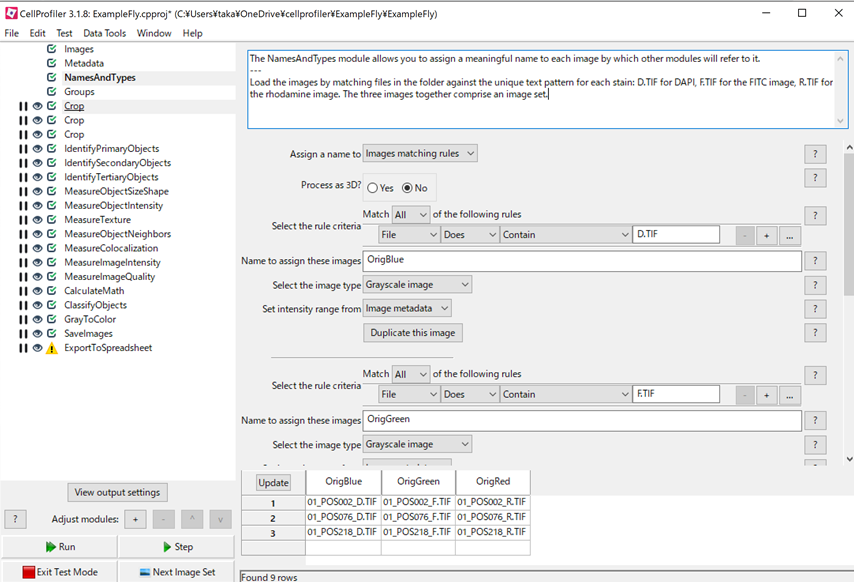
Cellprofiler Output During Processing Rightfeti Using cellprofiler and fcs express 4 image cytometry: joint webinar with denovo software on using cellprofiler to peform basic image assays and generate data compatible with export into fcs express. The steps for exporting data from cellprofiler have been outlined by illustrating how to add images, extract metadata from files, and modify the required modules according to the section1pipeline.cpproj. When specified, cellprofiler will output one line to the terminal per job to be run. this output should be further processed to generate a script that can invoke the jobs in a cluster computing context. Cellprofiler can be used to analyze the resulting images from imaging flow cytometry, whether brightfield, darkfield, or fluorescence. the ifc website page has further details on this workflow.
Comments are closed.