Building A Genome Alignment Visualization Tool In Python Peerdh

Building A Genome Alignment Visualization Tool In Python Peerdh Creating a genome alignment visualization tool can be a rewarding project. it allows you to visualize how different sequences align, making it easier to understand the relationships between them. this article will guide you through the process of building such a tool using python. Pygenomeviz is a genome visualization python package for comparative genomics implemented based on matplotlib. this package is developed for the purpose of easily and beautifully plotting genomic features and sequence similarity comparison links between multiple genomes.

Benchmarking Genome Alignment Algorithms In Python Peerdh This package is developed for the purpose of easily and beautifully plotting genomic features and sequence similarity comparison links between multiple genomes. Pygenomeviz is a genome visualization python package for comparative genomics implemented based on matplotlib. this package is developed for the purpose of easily and beautifully plotting genomic features and sequence similarity comparison links between multiple genomes. In this article, an educational visualization tool for biological sequence alignment is presented, and the source code is released in order to encourage the collaborative power of open source software, with the expectation of further contributions from the community in the future. We also design a new visualization method for genome alignments in multiple alignment format to explore the pattern of within and between species variation. combining these visual representations with prime knowledge, ggmsa assists researchers in discovering msa and making decisions.
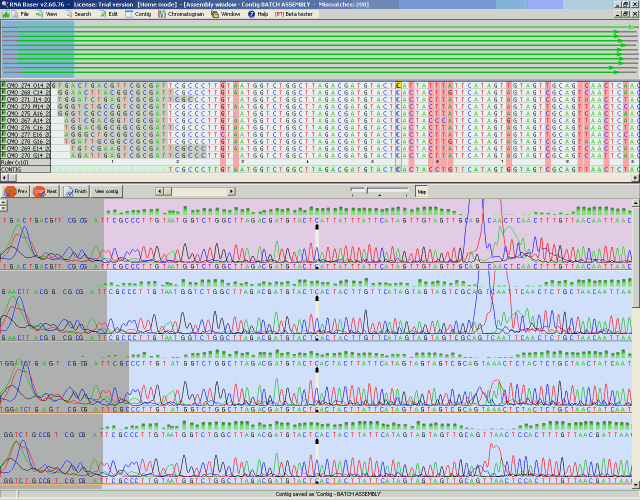
Building A Simple Sequence Alignment Tool Using Python Peerdh In this article, an educational visualization tool for biological sequence alignment is presented, and the source code is released in order to encourage the collaborative power of open source software, with the expectation of further contributions from the community in the future. We also design a new visualization method for genome alignments in multiple alignment format to explore the pattern of within and between species variation. combining these visual representations with prime knowledge, ggmsa assists researchers in discovering msa and making decisions. This package is developed for the purpose of easily and beautifully plotting genomic features and sequence similarity comparison links between multiple genomes. Proksee is an expert system for genome assembly, annotation and visualization, featuring interactive circular and linear genome maps. to begin using proksee, provide a complete genome sequence, sequencing reads or a cgview proksee map json file. Pairwise sequence alignment is the process of aligning two sequences to each other by optimizing the similarity score between them. It is able to create a genome graph using multiple whole genomes and existing multiple sequence alignment (msa) tools, allowing any current or future algorithm to be employed.

Building A Simple Sequence Alignment Tool Using Python Peerdh This package is developed for the purpose of easily and beautifully plotting genomic features and sequence similarity comparison links between multiple genomes. Proksee is an expert system for genome assembly, annotation and visualization, featuring interactive circular and linear genome maps. to begin using proksee, provide a complete genome sequence, sequencing reads or a cgview proksee map json file. Pairwise sequence alignment is the process of aligning two sequences to each other by optimizing the similarity score between them. It is able to create a genome graph using multiple whole genomes and existing multiple sequence alignment (msa) tools, allowing any current or future algorithm to be employed.
Building A Scoring System For Genome Alignment Algorithms In Python Pairwise sequence alignment is the process of aligning two sequences to each other by optimizing the similarity score between them. It is able to create a genome graph using multiple whole genomes and existing multiple sequence alignment (msa) tools, allowing any current or future algorithm to be employed.

Custom Scoring Functions For Genome Alignment Using Python Peerdh
Comments are closed.