Blast Tutorial 3 Docx Tutorial Blast Basic Local Alignment Search

Blast Basic Local Alignment Search Tool Pdf Sequence Alignment Enhanced document preview: tutorial: blast: basic local alignment search tool. the program uses heuristics to find and compare nucleotide or protein sequences to sequence databases and calculates the statistical significance of the matches. Mastering blast tutorial free download as word doc (.doc .docx), pdf file (.pdf), text file (.txt) or read online for free. blast (basic local alignment search tool) is a widely used bioinformatics program for comparing sequences to identify similarities and evolutionary relationships.

Blast Basic Local Alignment Search Tool Pdf Sequence Alignment The basic local alignment search tool (blast) finds regions of local similarity between sequences. the program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches. This week we will be looking at blast which is arguably the most successful and widely used bioinformatics algorithm ever. we show you how to use blast services at ncbi to do some practical sequence searching, investigating the possible functions and relationships between genes and proteins. Most often, local alignment ( “smith waterman”) is used for database searching: you are interested in finding out if any domain in your protein looks like something that is known. Global similarity algorithms optimize the overall alignment of two sequences, which may include large stretches of low similarity. 2. local similarity algorithms seek only relatively conserved sub sequences, and a single comparison may yield several distinct subsequence alignments.
Blast Basic Local Alignment Search Tool In Most often, local alignment ( “smith waterman”) is used for database searching: you are interested in finding out if any domain in your protein looks like something that is known. Global similarity algorithms optimize the overall alignment of two sequences, which may include large stretches of low similarity. 2. local similarity algorithms seek only relatively conserved sub sequences, and a single comparison may yield several distinct subsequence alignments. The document provides an introduction to blast (basic local alignment search tool), which is an algorithm used to compare gene and protein sequences to those in public databases. The basic local alignment search tool (blast) finds regions of local similarity between protein or nucleotide sequences. the program compares nucleotide or protein sequences to sequence in a database and calculates the statistical significance of the matches. Estimating similarities between biological sequences is becoming necessary to obtain hidden information present within the sequence and to trace evolutionary relationship exist within the. The basic local alignment search tool (blast) finds regions of local similarity between sequences. the program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches.
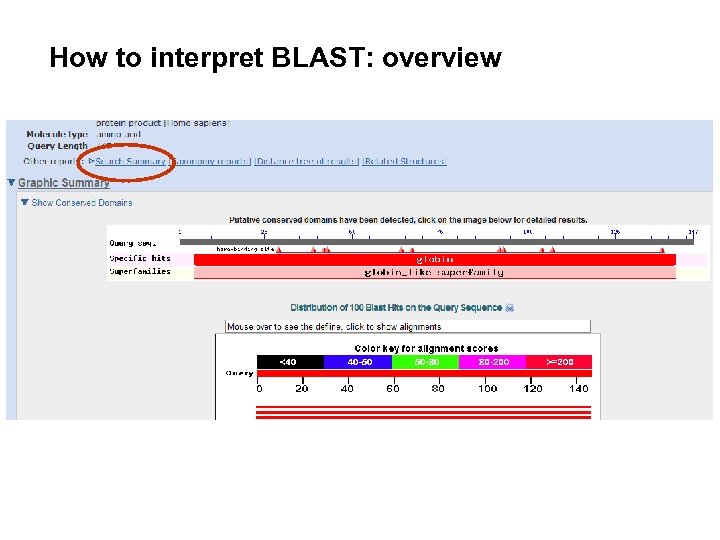

Blast Basic Local Alignment Search Tool In The document provides an introduction to blast (basic local alignment search tool), which is an algorithm used to compare gene and protein sequences to those in public databases. The basic local alignment search tool (blast) finds regions of local similarity between protein or nucleotide sequences. the program compares nucleotide or protein sequences to sequence in a database and calculates the statistical significance of the matches. Estimating similarities between biological sequences is becoming necessary to obtain hidden information present within the sequence and to trace evolutionary relationship exist within the. The basic local alignment search tool (blast) finds regions of local similarity between sequences. the program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches.

Blast Basic Local Alignment Search Tool In Estimating similarities between biological sequences is becoming necessary to obtain hidden information present within the sequence and to trace evolutionary relationship exist within the. The basic local alignment search tool (blast) finds regions of local similarity between sequences. the program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches.
Comments are closed.