Biox Nku
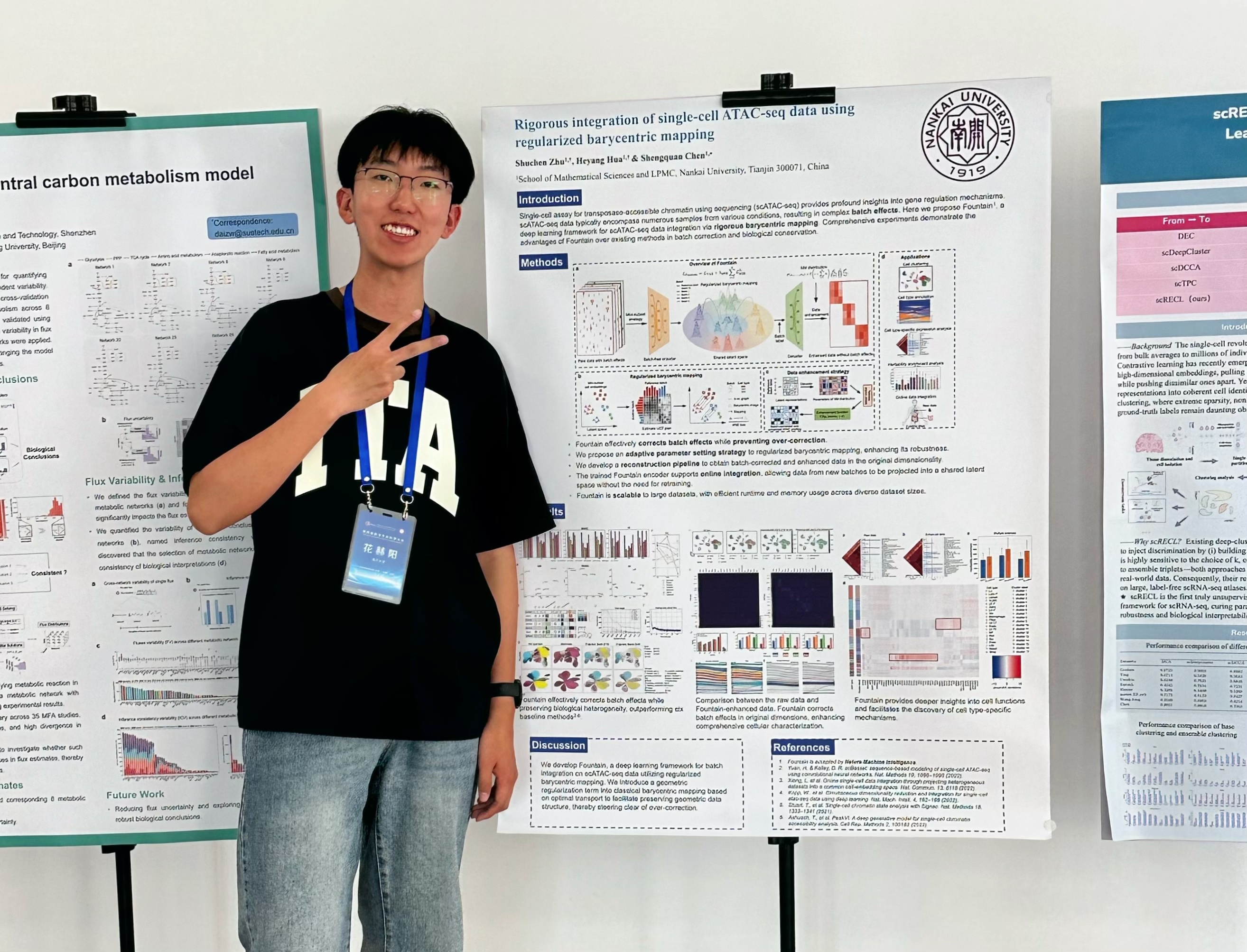
Biox Nku Join us now we are now looking for self motivated graduate undergraduate students. if you are interested in and want to join biox at nku, feel free to contact us. Biox nku has 28 repositories available. follow their code on github.
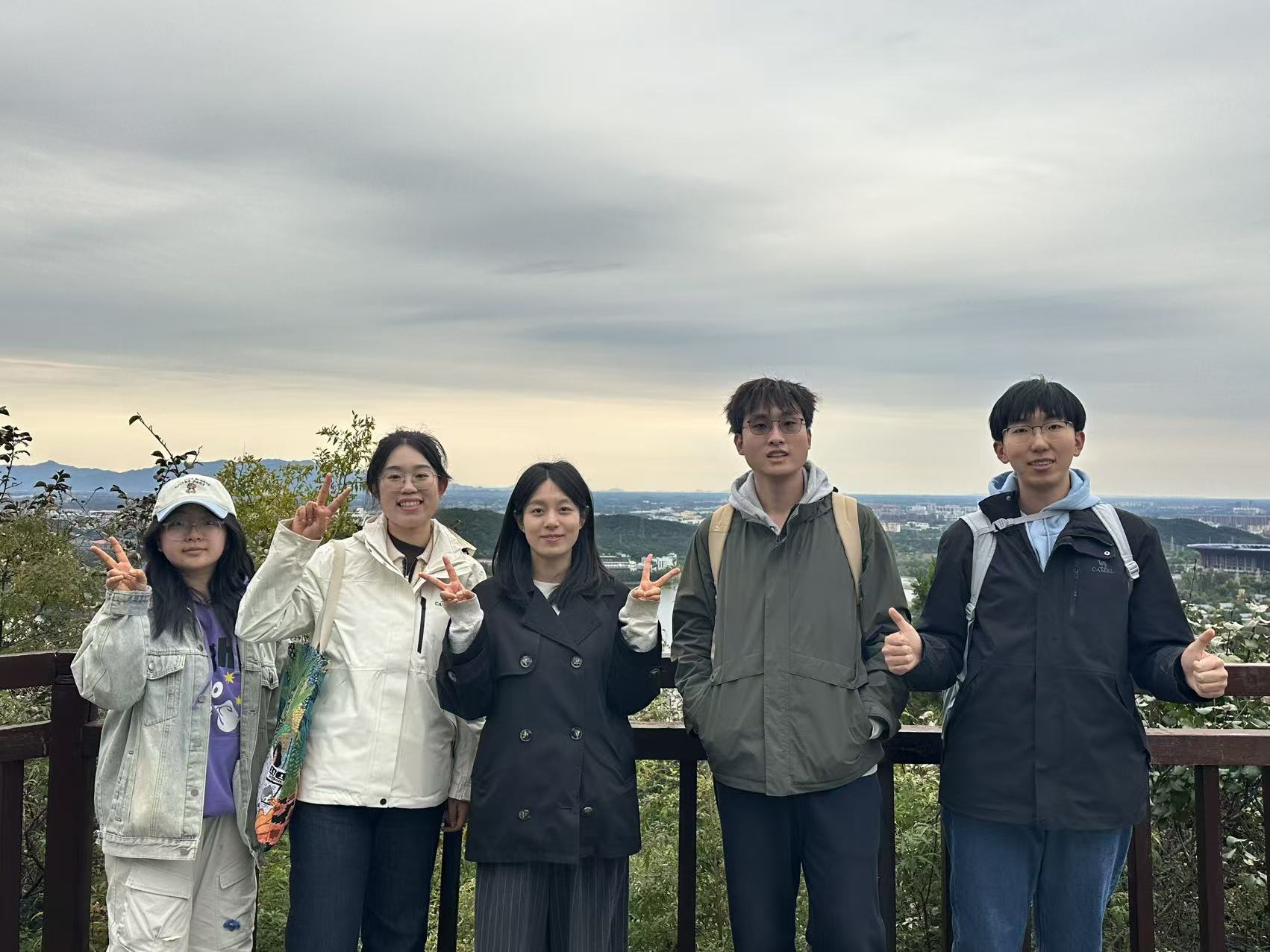

Biox Nku External resources archived in software heritage swh:1:dir:1cc55de47b78d7470d69bb78b2df2764789c4553 available in biox nku fountain release: v1.0.2 indexed in openaire. The epicarousel software is well documented and freely available at github biox nku epicarousel. it can be seamlessly interoperated with extensive sccas analysis toolkits. All the results demonstrate the superior performance of rainbow in cell type annotation for sccas data. we anticipate that rainbow will offer essential guidance and great assistance in sccas data analysis. the source codes are available at the github website (biox nku rainbow). It is a comprehensive dna methylation database that not only includes extensive dna methylation profiles across different ages and tissues but also encompasses aging related differentially methylated sites & regions.
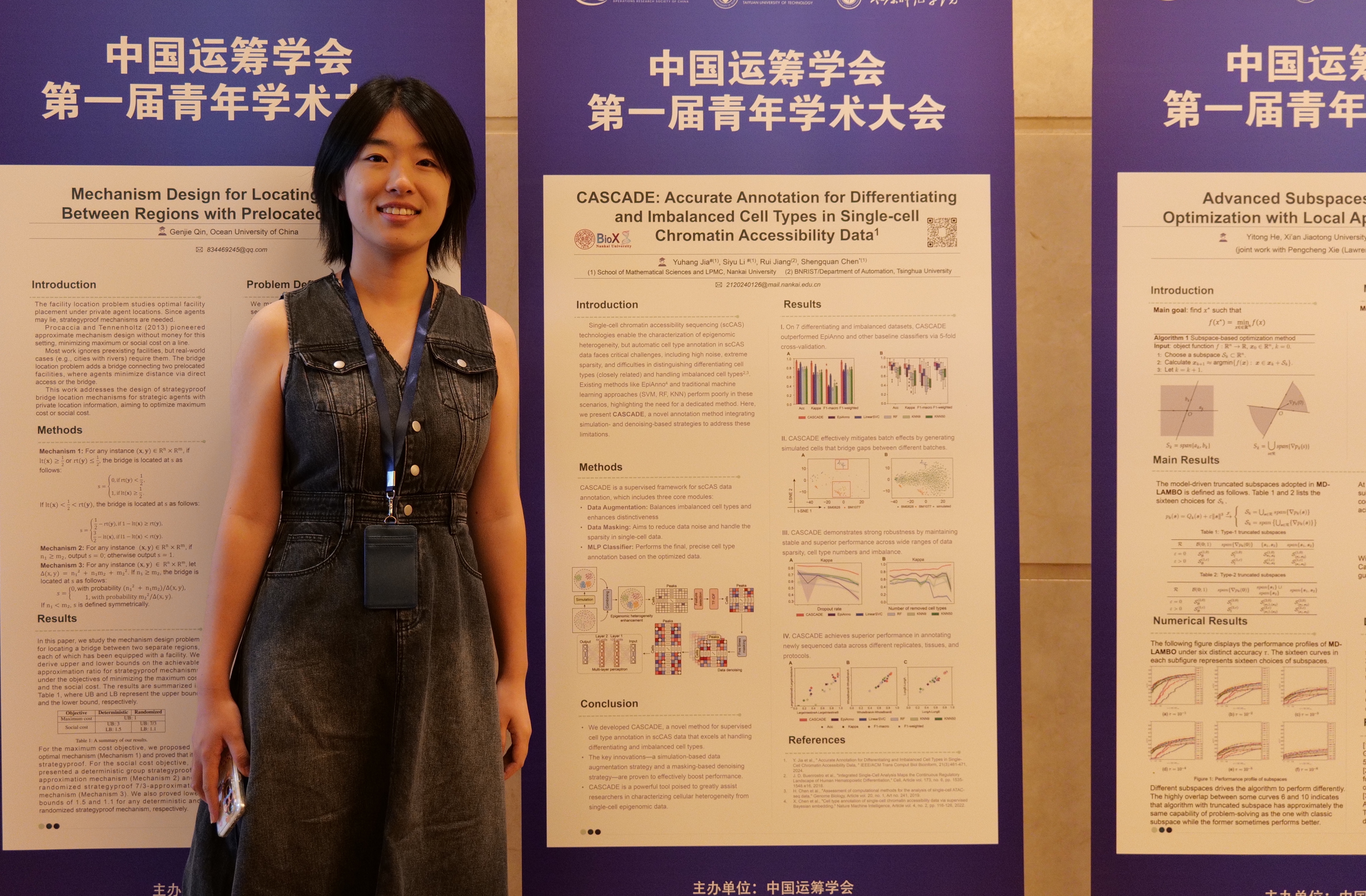
Biox Nku All the results demonstrate the superior performance of rainbow in cell type annotation for sccas data. we anticipate that rainbow will offer essential guidance and great assistance in sccas data analysis. the source codes are available at the github website (biox nku rainbow). It is a comprehensive dna methylation database that not only includes extensive dna methylation profiles across different ages and tissues but also encompasses aging related differentially methylated sites & regions. Estimate the number of cell types in sccas data by aster. – anndata object of shape n obs × n vars. rows correspond to cells and columns to genes. – list of optional numbers of cell types for the estimation. – whether to convert the count matrix into a binary matrix. by default, binary=true. Biox at nku about the company biox at nku is an unfunded company. it operates as a developer of mathematical and information technology for single cell omics data. biox at nku has not raised any funding yet. We provide a tutorial for running sccase and integrating episcanpy for sccas data analysis. the expected run time for the tutorial is less than 5 minutes. the extension of sccase can additionally correct sequencing depth, and we provide a tutorial. to enhance datasets with batch effects, refer to the tutorial. Aster: accurate estimation of cell type numbers in single cell chromatin accessibility data. aster provides an accurate and efficient way to estimate the number of cell types in single cell chromatin accessibility data.
Comments are closed.