Best Practices For Sequence Variant Analysis

Best Practices For Sequence Variant Analysis This document describes and discusses the various processes applied to high throughput sequencing data analysis with the intent of providing key best practices for germline variant analysis via sgs and tgs. In this review, we will discuss current approaches to variant calling using sequencing data and specifically focus on considerations while planning genomics experiments, practices while executing them, and promising developments to consider for future projects.
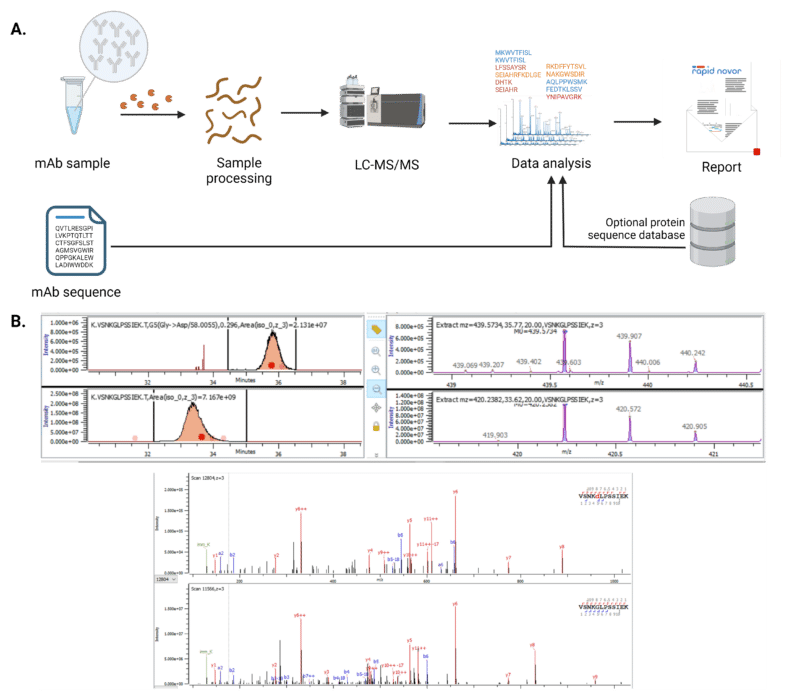
Sequence Variant Analysis Of Monoclonal Antibodies The 2019 casss workshop focused on emerging best practices and strategies for carrying out ms based sequence variant analysis (sva) via peptide mapping on recombinant therapeutic proteins. Roundtable 5 notes: mass spectrometry 2023 best practices for sequence variant analysis. This article provides a comprehensive overview of the best practices for variant calling in clinical sequencing, covering data preprocessing, alignment, variant detection algorithms, filtering, and validation. By following these best practices, the document aims to increase the diagnostic accuracy for hereditary diseases, facilitating early diagnosis, prevention, and personalized treatment strategies.
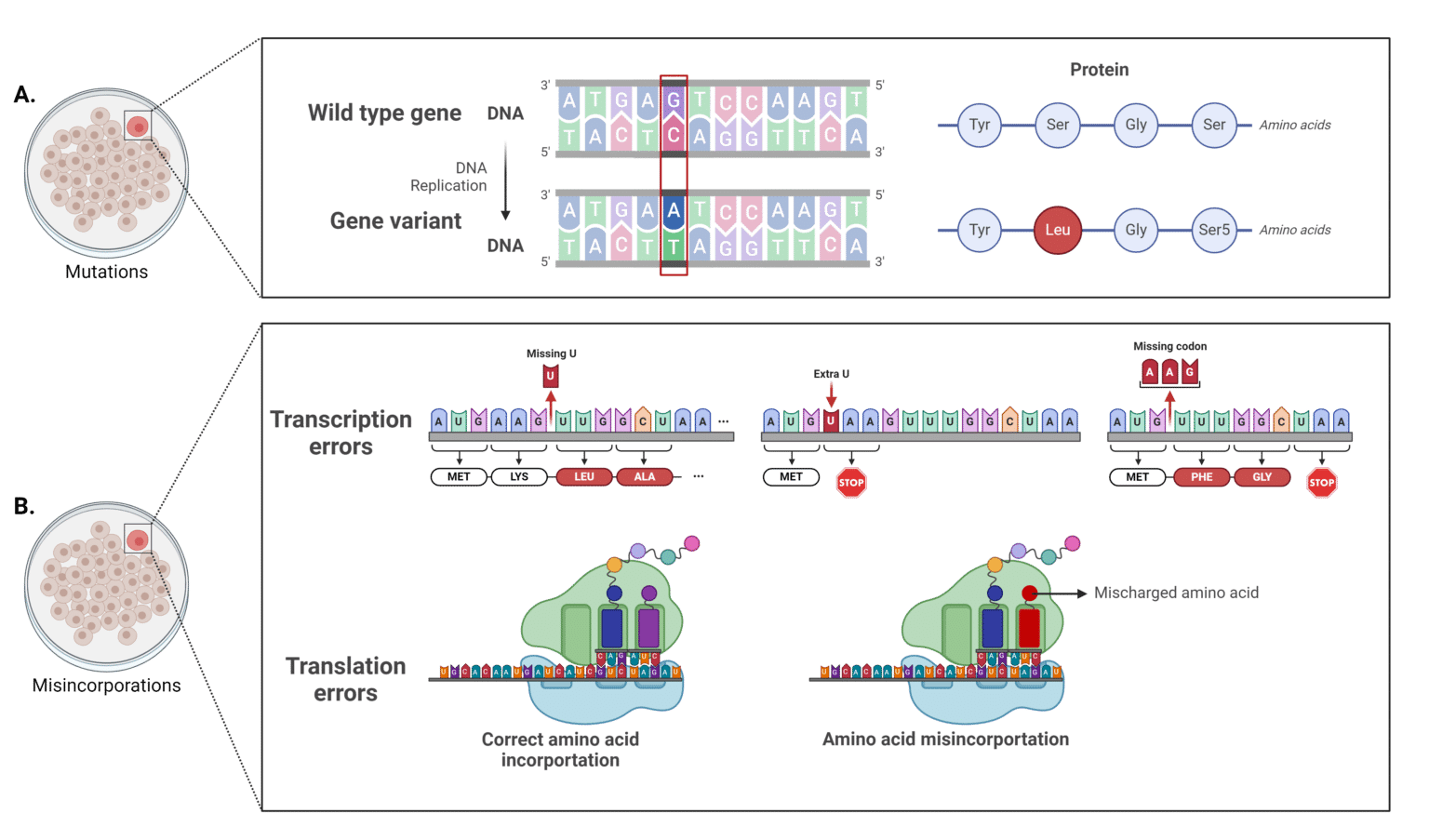
Sequence Variant Analysis Of Monoclonal Antibodies This article provides a comprehensive overview of the best practices for variant calling in clinical sequencing, covering data preprocessing, alignment, variant detection algorithms, filtering, and validation. By following these best practices, the document aims to increase the diagnostic accuracy for hereditary diseases, facilitating early diagnosis, prevention, and personalized treatment strategies. In this review, i discuss the current best practices for variant calling in clinical sequencing studies, with a particular emphasis on trio sequencing for inherited disorders and somatic. Tl;dr: this comprehensive review outlines best practices for analyzing germline variants and dna methylation using second and third generation sequencing data, emphasizing meticulous data handling, advanced methodologies, and bioinformatics tools to ensure accurate and reproducible genetic analyses. This comprehensive review provides insights and suggested strategies for the analysis of germline variants using second and third generation sequencing technologies (sgs and tgs). We use the genome analysis toolkit and the best practices for variant discovery analysis outlined by the broad institute. each of the steps in the flowchart below is explained within the step by step protocols that follow.

Best Practices For Variant Annotation And Filtration In this review, i discuss the current best practices for variant calling in clinical sequencing studies, with a particular emphasis on trio sequencing for inherited disorders and somatic. Tl;dr: this comprehensive review outlines best practices for analyzing germline variants and dna methylation using second and third generation sequencing data, emphasizing meticulous data handling, advanced methodologies, and bioinformatics tools to ensure accurate and reproducible genetic analyses. This comprehensive review provides insights and suggested strategies for the analysis of germline variants using second and third generation sequencing technologies (sgs and tgs). We use the genome analysis toolkit and the best practices for variant discovery analysis outlined by the broad institute. each of the steps in the flowchart below is explained within the step by step protocols that follow.

Best Practices For Variant Annotation And Filtration This comprehensive review provides insights and suggested strategies for the analysis of germline variants using second and third generation sequencing technologies (sgs and tgs). We use the genome analysis toolkit and the best practices for variant discovery analysis outlined by the broad institute. each of the steps in the flowchart below is explained within the step by step protocols that follow.
Comments are closed.