Bcftools Variant Statistics
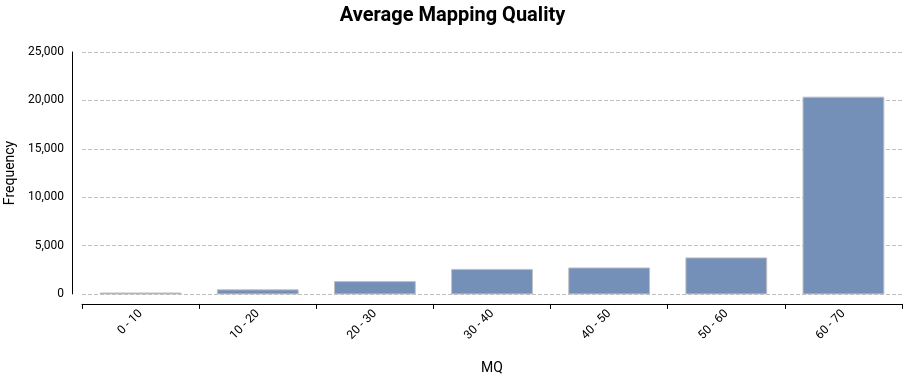
Variant Calling Using Bcftools Omicsbox User Manual Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed.
Variant Calling Using Bcftools Omicsbox User Manual The stats command (vcfstats.c) provides comprehensive statistical analysis of vcf bcf files, generating detailed reports on variant composition, quality metrics, and sample level statistics. Bcftools is a suite of utilities designed for manipulating and analyzing variant call data stored in vcf and bcf formats. it provides essential tools for filtering, converting, and summarizing vcf information, making it an important component of most variant calling workflows. Bcftools stats: this is a command from the bcftools suite used to generate statistics about variants in a vcf file. it calculates various statistics, including the number of reference homozygous (nrefhom), non reference homozygous (nnonrefhom), heterozygous (nhets), and other metrics. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed.
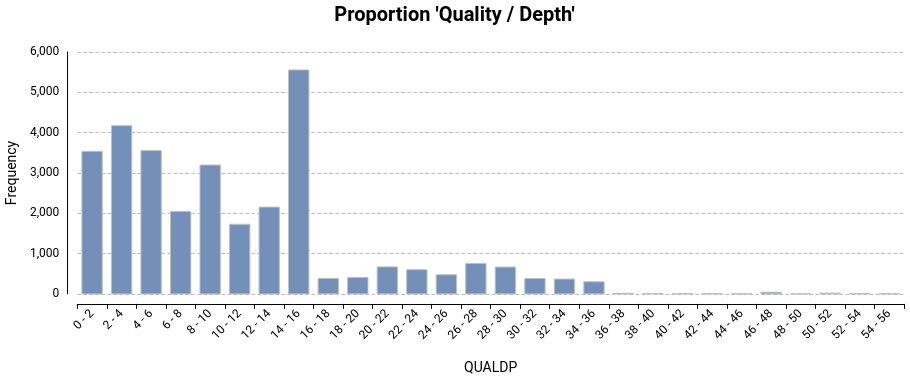
Variant Calling Using Bcftools Omicsbox User Manual Bcftools stats: this is a command from the bcftools suite used to generate statistics about variants in a vcf file. it calculates various statistics, including the number of reference homozygous (nrefhom), non reference homozygous (nnonrefhom), heterozygous (nhets), and other metrics. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed. Bcftools is a suit of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. the tool is designed for handling data streams and as such treats input output files as stdin and stdout. Let's use this data now to generate some variant and genotype calls inside fut2. conceptually genotype calling with bcftools has two steps: the first step looks at each sample seperately and compute genotype likelihoods for each possible genotype at each site. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed.
Variant Calling Using Bcftools Omicsbox User Manual Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed. Bcftools is a suit of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. the tool is designed for handling data streams and as such treats input output files as stdin and stdout. Let's use this data now to generate some variant and genotype calls inside fut2. conceptually genotype calling with bcftools has two steps: the first step looks at each sample seperately and compute genotype likelihoods for each possible genotype at each site. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed.
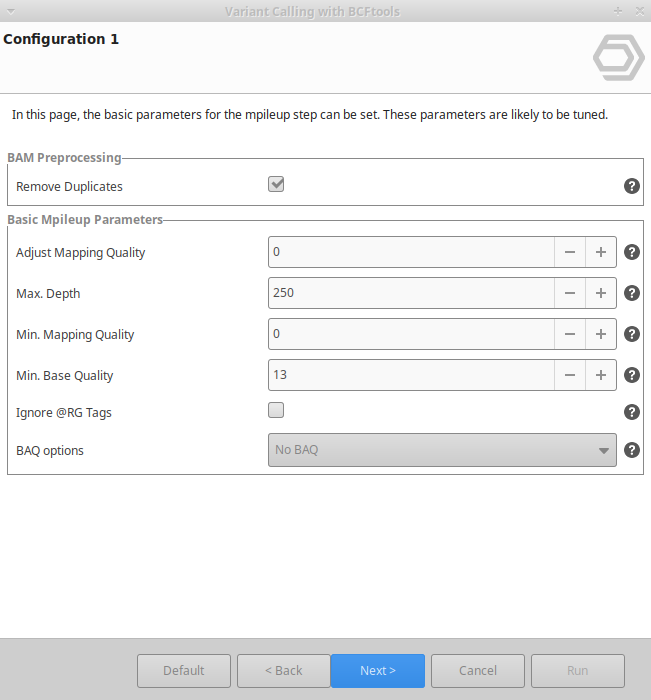
Variant Calling Using Bcftools Omicsbox User Manual Let's use this data now to generate some variant and genotype calls inside fut2. conceptually genotype calling with bcftools has two steps: the first step looks at each sample seperately and compute genotype likelihoods for each possible genotype at each site. Bcftools is a set of utilities that manipulate variant calls in the variant call format (vcf) and its binary counterpart bcf. all commands work transparently with both vcfs and bcfs, both uncompressed and bgzf compressed.
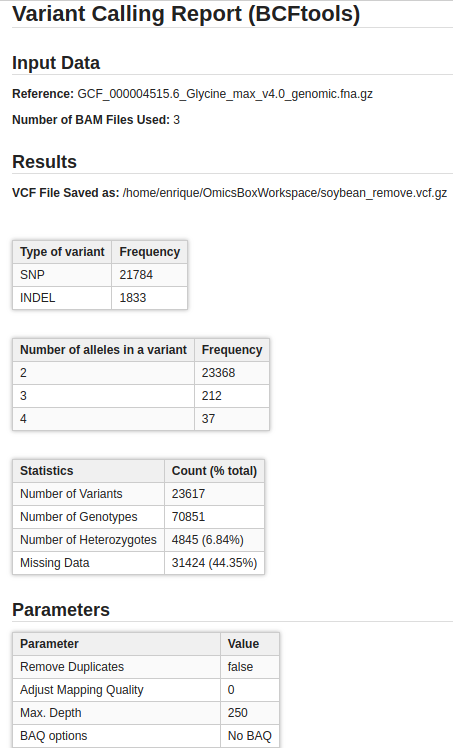
Variant Calling Using Bcftools Omicsbox User Manual
Comments are closed.