5 9 Space Efficient Sequence Alignment
Github Jiazheng Yuan Sequence Alignment Space Efficient Iterative Enjoy the videos and music you love, upload original content, and share it all with friends, family, and the world on . Learn how to perform sequence alignments using linear space algorithms to efficiently compare biological sequences with reduced memory usage.

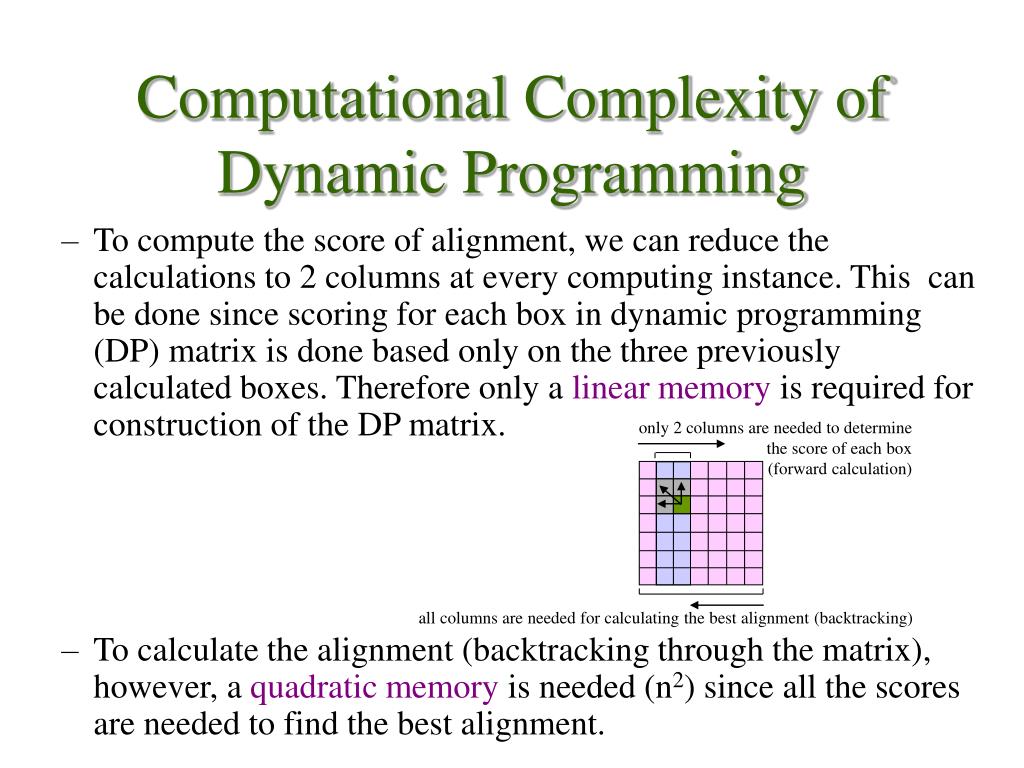
Sequence Alignment Assignment Point The space efficient sequence alignment algorithm uses only two columns every time to calculate the value of the optimal alignment score. we keep copying the current column values to the previous column and update the current column with the new values. Saul b. needleman and christian d. wunsch devised a dynamic programming algorithm to the problem and got it published in 1970. since then, numerous improvements have been made to improve the time complexity and space complexity, however these are beyond the scope of discussion in this post. Alignment score • space complexity of computing just the score itself is o(n) we only need the previous column to calculate the current column, and we can then throw away that previous column once we’re done using it. Best alignment between prefix of x and y score of an optimal alignment between y and a prefix of x c 9.
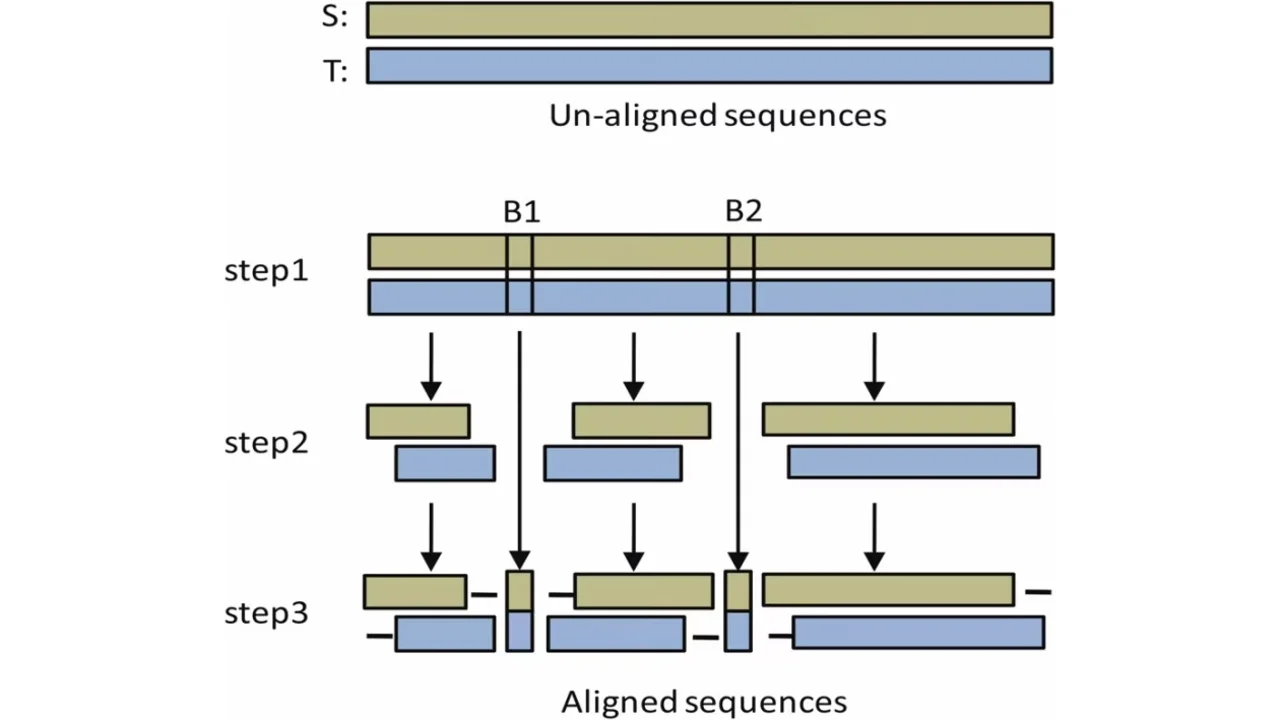
Efficient Sequence Alignment Technique For Bioinformatics Sensors Alignment score • space complexity of computing just the score itself is o(n) we only need the previous column to calculate the current column, and we can then throw away that previous column once we’re done using it. Best alignment between prefix of x and y score of an optimal alignment between y and a prefix of x c 9. Trivially, we can compute the cost of an optimal alignment in linear space by arranging subproblems intelligently we can define a “reverse” dp that works on suffixes instead of prefixes combining the “forward” and “reverse” dp using a divide and conquer technique, we can compute the optimal solution. The sequence alignment problem is fundamental in bioinformatics; we have implemented the $x$ drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the. The sequence alignment problem is fundamental in bioinformatics; we have implemented the x drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the graphcore intelligence processor unit (ipu) accelerator. Quicked’s library, based on the bound and align strategy, offers a practical and efficient method for approximate and exact sequence alignment of long and noisy sequences.

Ppt Space Efficient Sequence Alignment Techniques In Bioinformatics Trivially, we can compute the cost of an optimal alignment in linear space by arranging subproblems intelligently we can define a “reverse” dp that works on suffixes instead of prefixes combining the “forward” and “reverse” dp using a divide and conquer technique, we can compute the optimal solution. The sequence alignment problem is fundamental in bioinformatics; we have implemented the $x$ drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the. The sequence alignment problem is fundamental in bioinformatics; we have implemented the x drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the graphcore intelligence processor unit (ipu) accelerator. Quicked’s library, based on the bound and align strategy, offers a practical and efficient method for approximate and exact sequence alignment of long and noisy sequences.

Ppt Space Efficient Sequence Alignment Techniques In Bioinformatics The sequence alignment problem is fundamental in bioinformatics; we have implemented the x drop algorithm, a heuristic method for pairwise alignment that reduces search space, on the graphcore intelligence processor unit (ipu) accelerator. Quicked’s library, based on the bound and align strategy, offers a practical and efficient method for approximate and exact sequence alignment of long and noisy sequences.
Comments are closed.