2025 Page 2 Mixomics

Home Mixomics Pro You can install the latest version of mixomics on bioconductor here. this latest release version of the package runs on r version 4.4 and includes some minor bug fixes and updated code and unit tests. This version of mixomics is a pre release which includes changes to the performance assessment and parameter tuning workflow, to be shared so this new workflow can be beta tested.

2025 Page 2 Mixomics Mixomics is an r toolkit dedicated to the exploration and integration of biological data sets with a specific focus on variable selection. the package currently includes more than twenty multivariate methodologies, mostly developed by the mixomics team (see some of our references in 1.2.3). The data that can be analysed with mixomics may come from high through put sequencing technologies, such as omics data (transcriptomics, metabolomics, pro teomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imag ing). Here is an overview of the most widely used methods in mixomics that will be further detailed in this vignette, with the exception of rcca. we depict them along with the type of data set they can handle. Beginners looking for an introduction to mixomics methods for single – and multi omics analyses. current mixomics users who want to deepen their understanding of the mixomics methods.
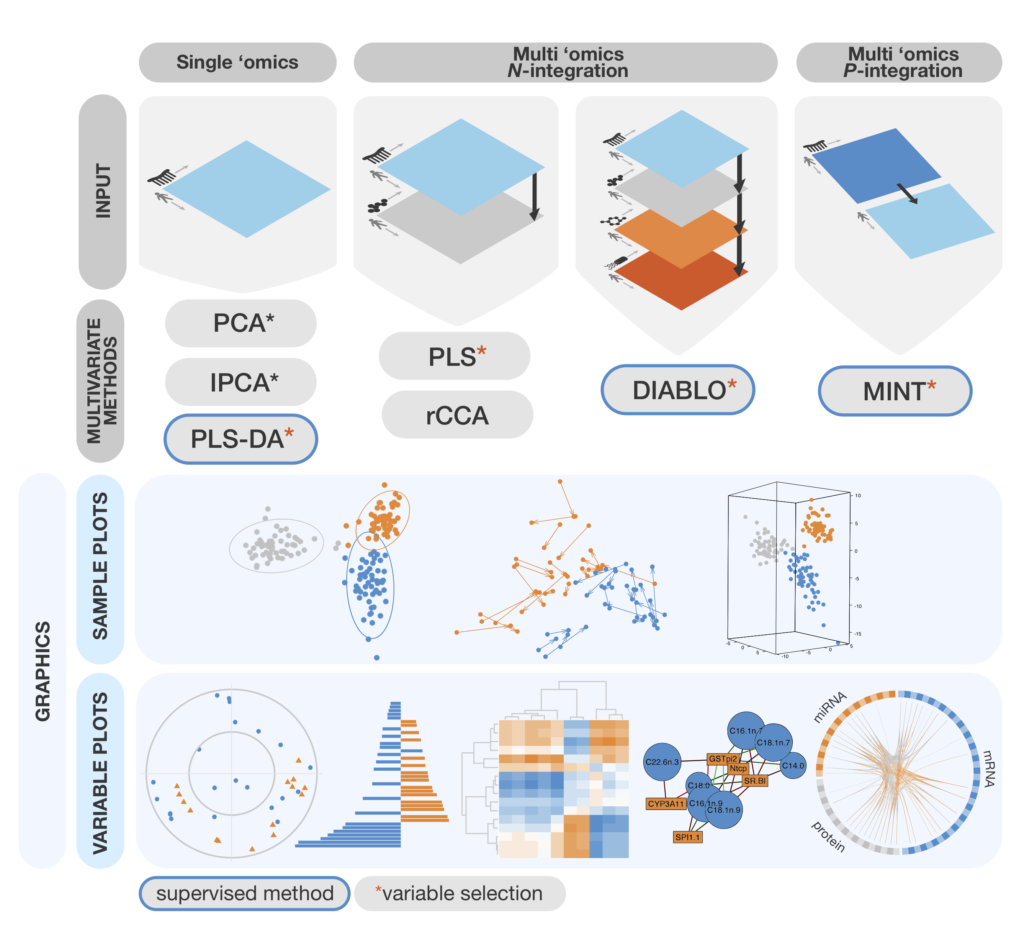
Mixomics Omics Data Integration Project Here is an overview of the most widely used methods in mixomics that will be further detailed in this vignette, with the exception of rcca. we depict them along with the type of data set they can handle. Beginners looking for an introduction to mixomics methods for single – and multi omics analyses. current mixomics users who want to deepen their understanding of the mixomics methods. Bioconductor versions are updated twice a year, between these updates you can download the latest version of mixomics from github. note that this latest version of mixomics is under development and may not be stable, check the github page for releases which have passed package testing. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). Beginners looking for an introduction to mixomics methods for single and multi omics analyses. current mixomics users who want to deepen their understanding of the mixomics methods. You can install our latest stable github version of mixomics via our docker container. you can do this by downloading and using the docker desktop application or via the command line as described below.

Collaborations Mixomics Bioconductor versions are updated twice a year, between these updates you can download the latest version of mixomics from github. note that this latest version of mixomics is under development and may not be stable, check the github page for releases which have passed package testing. The data that can be analysed with mixomics may come from high throughput sequencing technologies, such as omics data (transcriptomics, metabolomics, proteomics, metagenomics etc) but also beyond the realm of omics (e.g. spectral imaging). Beginners looking for an introduction to mixomics methods for single and multi omics analyses. current mixomics users who want to deepen their understanding of the mixomics methods. You can install our latest stable github version of mixomics via our docker container. you can do this by downloading and using the docker desktop application or via the command line as described below.

Methods Mixomics Beginners looking for an introduction to mixomics methods for single and multi omics analyses. current mixomics users who want to deepen their understanding of the mixomics methods. You can install our latest stable github version of mixomics via our docker container. you can do this by downloading and using the docker desktop application or via the command line as described below.

Open 3 Day Mixomics Workshop In Person 22 24 Sept 2025 Lund
Comments are closed.